We are often asked if it’s possible to work with ontologies in KNIME Analytics Platform.
With “work with ontologies” people can mean many different things but let's focus today on one particular ontology and basic tasks including reading and querying ontologies to create an interactive tool at the end. For this purpose, today, we dive into the world of chemistry to use the ChEBI ontology (Chemical Entities of Biological Interest).
Even if chemistry is not a domain of interest for you, this blog post can still be of high value as we, for example, show how to read an OWL file, how to create queries in SPARQL as well as different possibilities for visualizing ontology content in an interactive composite view. How you adopt this to your own use case, ontology, and extracted dataset, we will leave to you and your imagination.

ChEBI
Especially in the Life Sciences area, ontologies are very popular and frequently used for different purposes such as data integration, curation, defining standards, or labeling. Just how important ontologies are is illustrated by facts that sources like the BioPortal 1 contain already more than 800 ontologies in their repositories.
With the workflow described in this blog post, we will demonstrate a way to explore ChEBI2 , which is a freely available ontology containing a classification of chemical compounds as well as information about the role of compounds like their application or the biological and chemical role. It contains in general three main classifications like chemical entity, role, and subatomic particle. In this workflow, we use molecular entity and role class (See Fig. 1).
ChEBI can be downloaded in different file formats - today we will work with an OWL file which can be downloaded here.
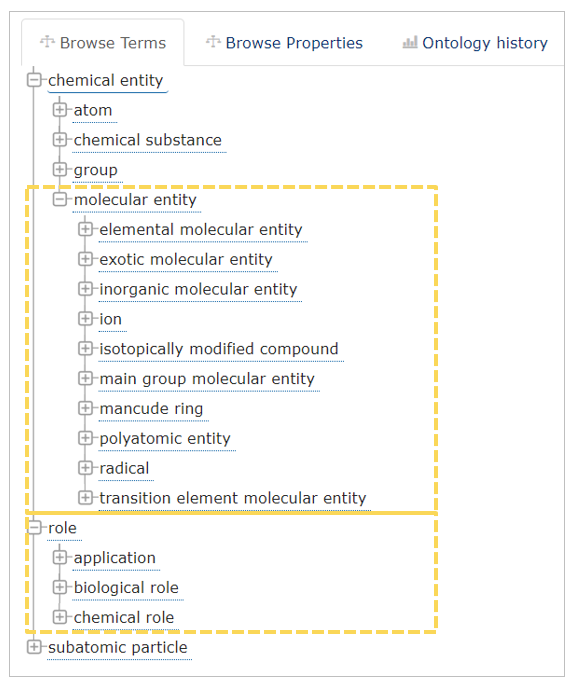
Fig. 1: Overview of ChEBI classes (screenshot taken from here). The yellow boxes show which parts will be used and explored in the workflow. The remaining parts are ignored.
Let’s start!
How to basically read and query ontologies stored in the OWL format is described in the blog post Will They Blend? KNIME Meets the Semantic Web. In the example described in this article, we used a pizza ontology to show how easy it is to explore that type of data.
With the following example workflow, we play with the terms and content of the ChEBI ontology while combining searches, results and data in order to create interactive views where the content can be explored. We hope to learn about compounds, their biological and chemical roles, as well as definitions and other sources that contain references to a particular compound.
This analysis was realized in the workflow depicted in Fig. 2 and contains the following main steps:
Step 1. Reading the OWL file into a SPARQL Endpoint
Step 2. Substructure search & selecting a role of a chemical compound
Step 3. View compounds matching the substructure search and role. Here one compound needs to be selected
Step 4. Show the selected compound in a network with all their parent classes, hierarchies and roles. Select a disease in the Tag Cloud to merge some more data in the next step
Step 5. Viewing results from selection in Step 4
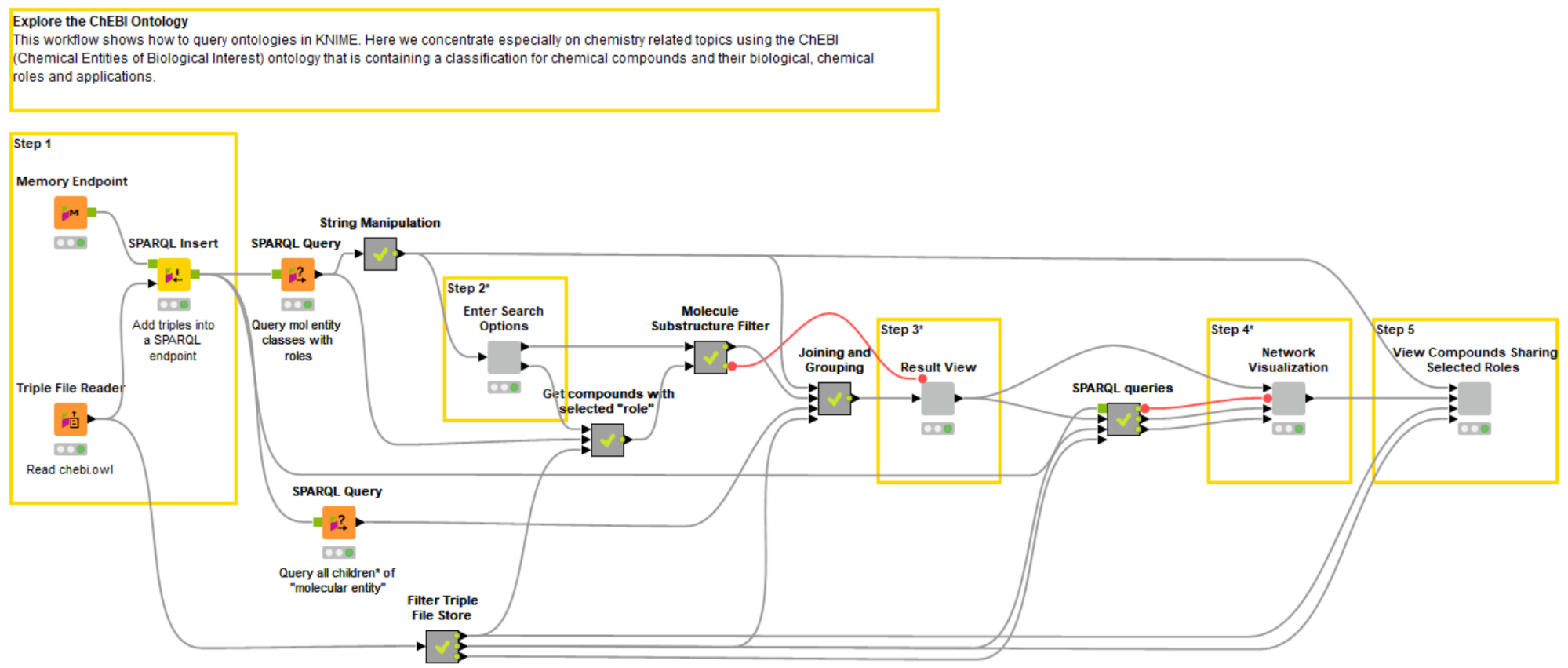
Fig. 2: Example workflow showing how to explore the ChEBI ontology stored in OWL format.
Step 1. Reading the OWL file into a SPARQL Endpoint
Analogous to the previously described use case of a pizza ontology, we use the Triple File Reader node to read the OWL file and insert the list of triples into a SPARQL Endpoint which is connected to a Memory Endpoint Node (See Fig. 2, Step 1). With this in place, and successfully executed, we now have the basis to start writing and executing SPARQL queries as well as filtering information from the list of Triples.
Quick reminder here: RDF triple - also known as semantic triple - always contains three columns: subject, predicate, and object. Read more here.
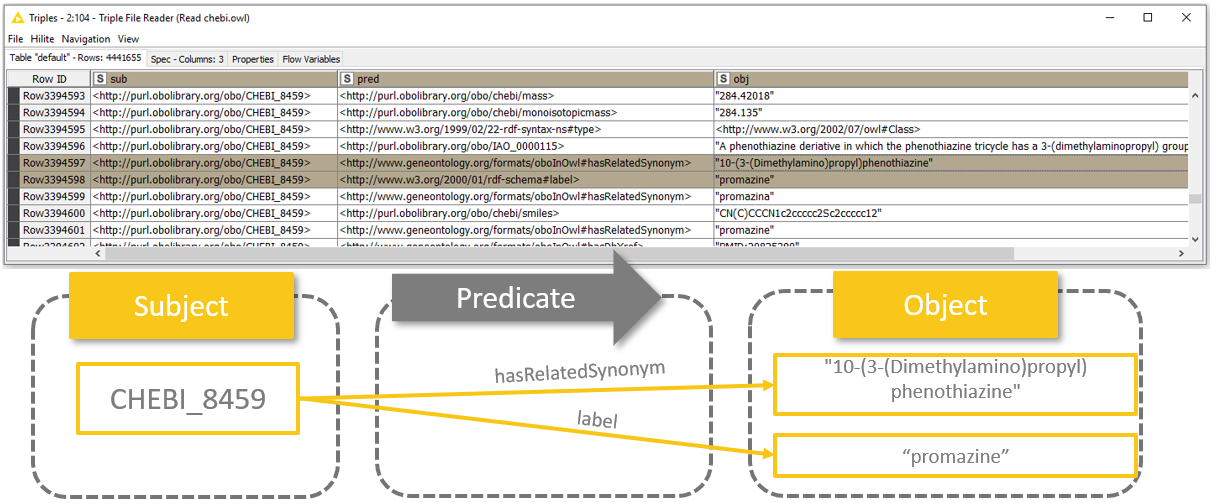
Fig. 3: Schema showing how to interpret RDF triples.
Step 2. Substructure search paired with role information
Imagine a scientist is investigating a new compound in development. She knows the chemical structure and the application of that compound but is curious to see if there are other compounds in ChEBI with similar properties. In this example workflow a SMILES for a substructure search can be added and the application or biological/chemical role of the compound can be selected (see Fig. 4).
Therefore, the “Enter Search Options” component will be used to create a search query in order to add the above mentioned properties (Right click the component → select interactive view).
To allow insertion of a SMILES, we have used the String Widget node. Here, in Fig. 4., we added a Phenothiazine.
Let’s select a dopaminergic antagonist as the role. This is frequently used in anti-psychotic drugs for treating schizophrenia, bipolar disorders or psychosis stimulants.
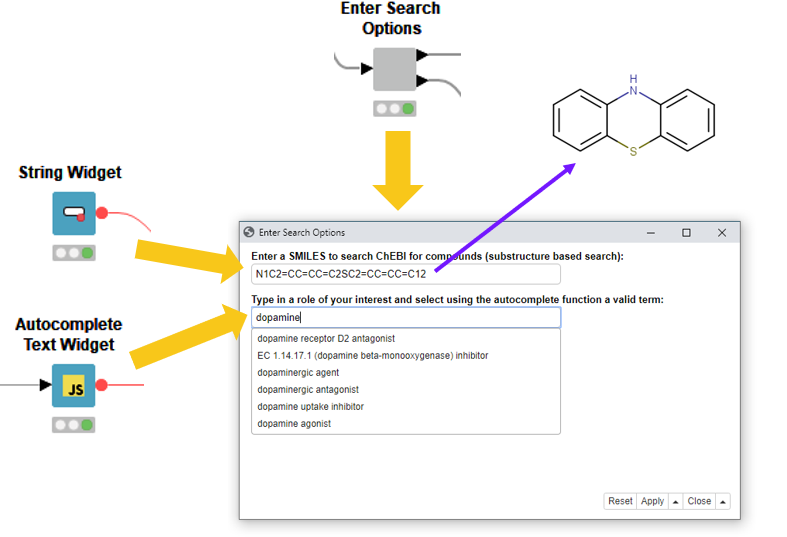
Fig. 4: The “Enter Search Options” component contains two options to enter input for a substructure search as well as a search for compounds; a specific role is assigned in the ontology.
Little hint here: As an alternative to the String Widget, a Molecule String Input node could also be used. This would give you the opportunity to draw a chemical structure instead of pasting a SMILES string.
Step 3. Let’s view first results!
In the following view (“Result View” component in the example workflow) you can inspect the results of the substructure search. Here the Tile View and RDKit Highlighting were used to show all compounds matching the entered search options. All these compounds are dopaminergic antagonists as selected in the previous view.
In this example we selected a compound called Promazine to see more information.
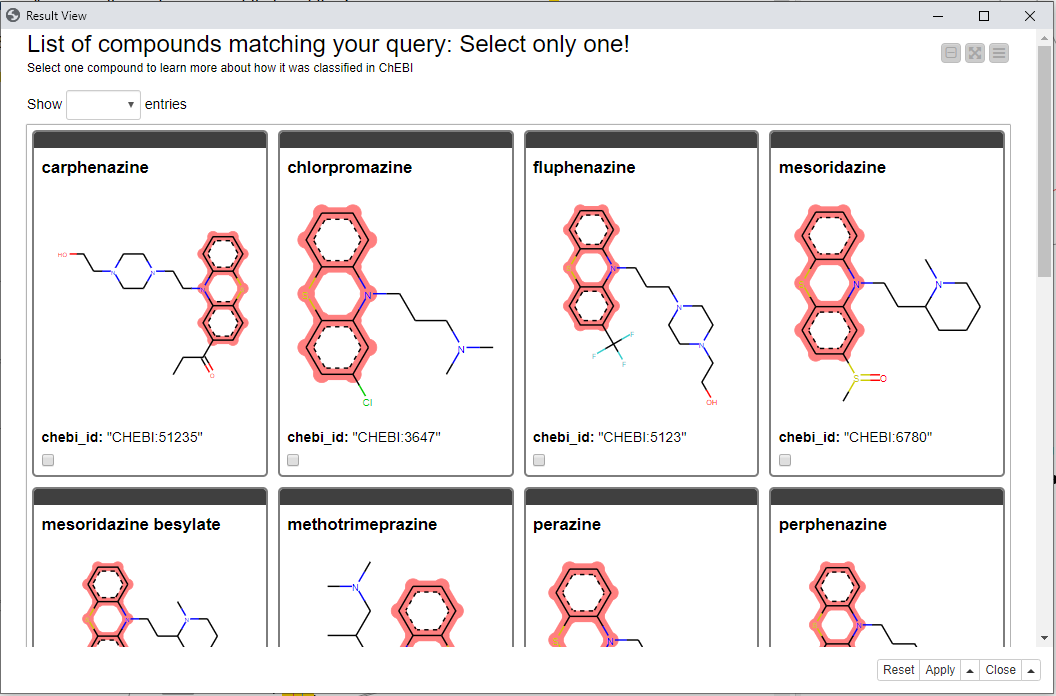
Fig. 5: Result view showing chemical compounds with the highlighted substructure using the Tile View node.
Step 4. Show a network with class information
In this example workflow, we also would like to show a way to visualize an ontology as a network. We do this with the Network Mining Extension. The view in this step is also interactive, which means whenever a node in the network is selected, the table under the network will show more information for the extracted entity, such as the definition, for example.
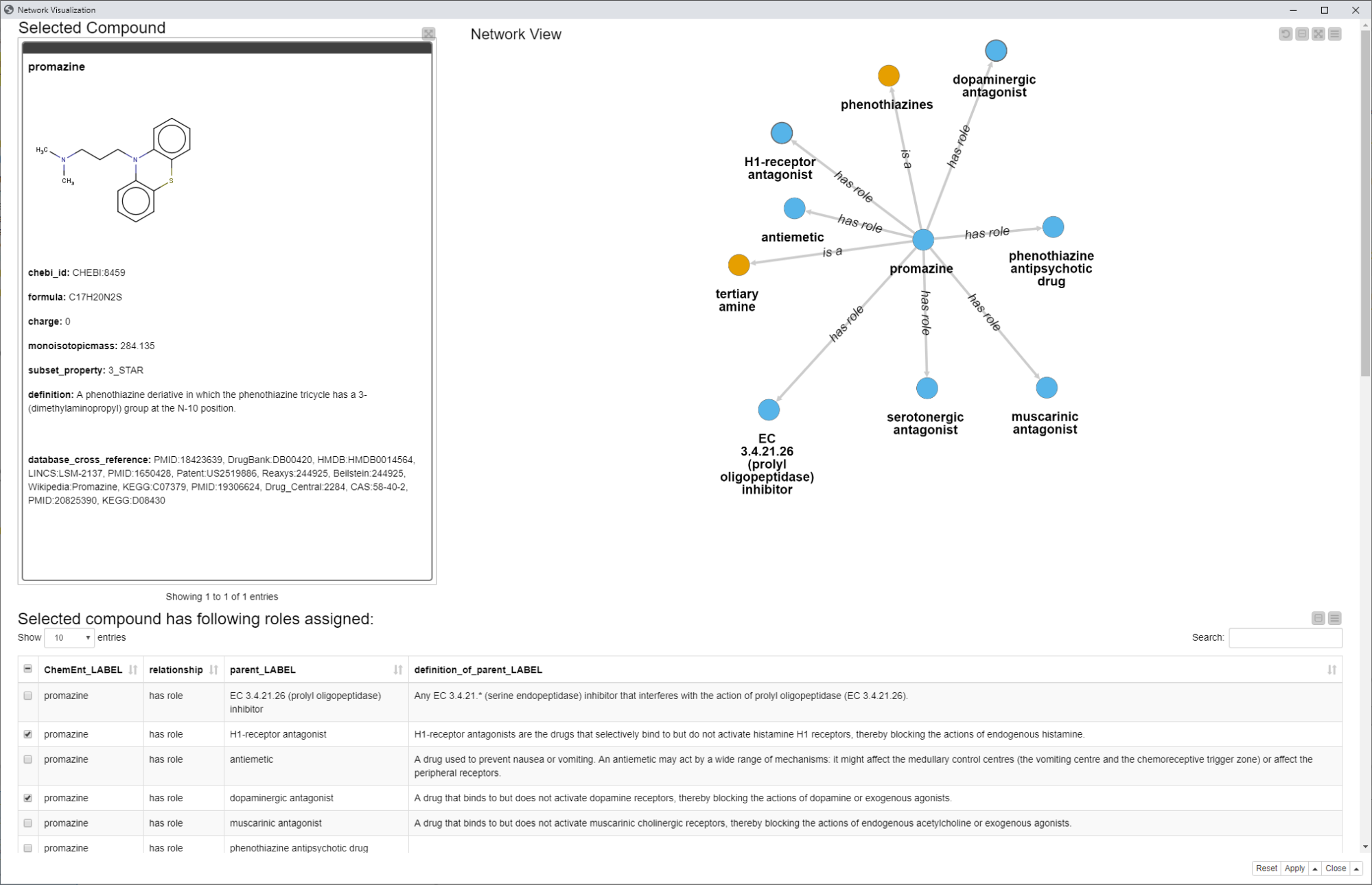
Fig. 6: Network view showing the selected compound as well as a network including the subClassOf connections as “has role” and “is a”. Additionally, more information such as definitions and synonyms can be selected and made visible for a node in the network.
If you scroll down, a second network becomes visible. This network contains entities starting from the selected compound, here Promazine, and shows how it has been classified in ChEBI. It shows all “is a” links from the compound through to the chemical entity. Additionally the “has role” links were added to show also here which roles are linked (blue nodes).
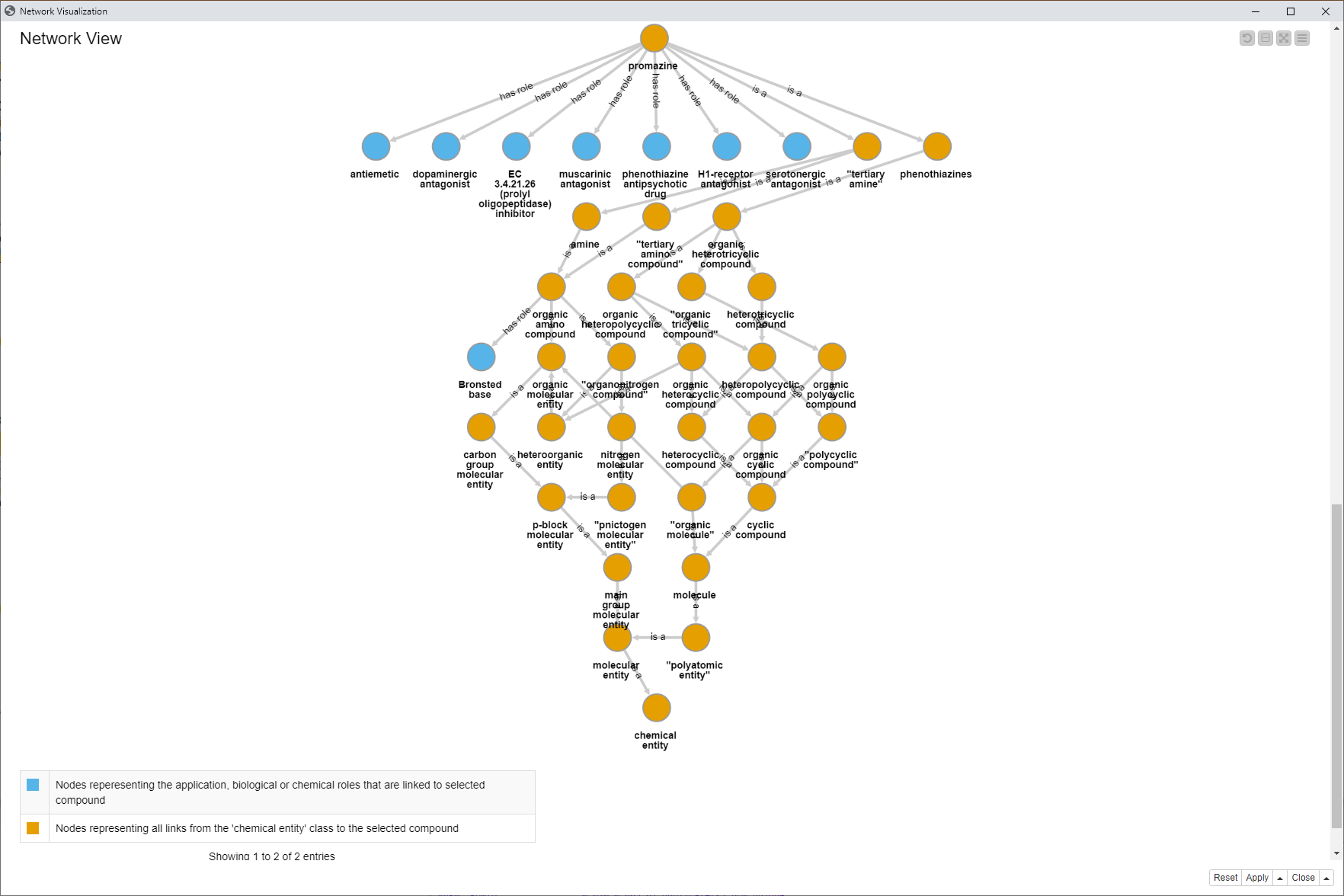
Fig. 7: Network view showing the selected compound as well as a network including the subClassOf connections as “has role” and “is a”.
Step 5. Show compounds sharing two roles
By looking at the network view in Fig. 6 and investigating the roles of a compound - let’s say you can now see another interesting role and you would like to see more compounds with that role - you can go one step further and select two different roles from the table like for example the already known dopaminergic antagonist in combination with the H1-receptor antagonist that plays a role in relieving allergic reactions.
In the last component “View Compounds Sharing Selected Roles” (Step 5) we see all those compounds containing both selected roles and the additional information, such as definitions, synonyms, or references to other ontologies, databases, and sources.
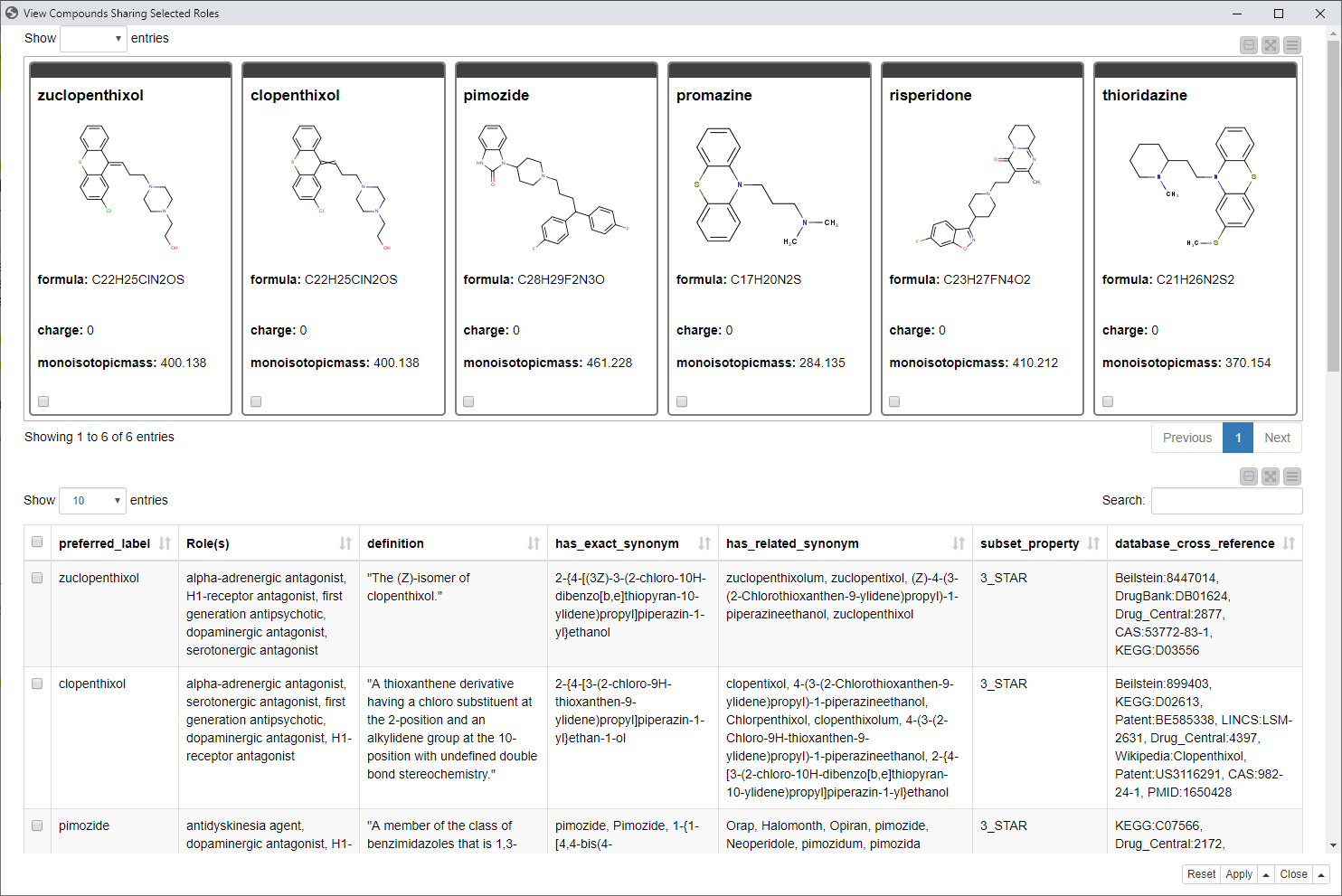
Fig. 8: Last interactive view of the workflow showing compounds with additional information having both selected “roles” from the network view.
Extensions needed to run the workflow:
- KNIME Semantic Web/Linked Data. To read more about that extension, you might find this blog article Integrating One More Data Source: The Semantic Web interesting, too.
- RDKit KNIME Integration is used to perform substructure searches and highlighting
- Network Mining Extension is used to create a network view
Instructions on how to install extensions can be found here.
Wrapping up
We started with an OWL file containing the ChEBI ontology and went through different steps of data exploration and visualization. We showed how to read an OWL file, how to create queries in SPARQL and presented different possibilities for visualizing ontology content in an interactive composite view. With this, we learned about the Promazine compound and the biological and chemical roles of that compound. We also discovered more about similar compounds and their roles, definitions, and synonyms.
The resulting data extracted from the ChEBI ontology can be directly explored using KNIME Analytics Platform. The workflow can also be deployed to KNIME Server where an expert of a certain research domain who is maybe not a KNIME or an ontology expert can analyze the data in the WebPortal without the need to write SPARQL queries.
The workflow described in this blog post, the ChEBI Ontology Explorer can be downloaded here from the KNIME Hub.
References
1. Whetzel PL, Noy NF, Shah NH, Alexander PR, Nyulas C, Tudorache T, Musen MA. BioPortal: enhanced functionality via new Web services from the National Center for Biomedical Ontology to access and use ontologies in software applications. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W541-5. Epub 2011 Jun 14.
2. Hastings J, Owen G, Dekker A, Ennis M, Kale N, Muthukrishnan V, Turner S, Swainston N, Mendes P, Steinbeck C. (2016). ChEBI in 2016: Improved services and an expanding collection of metabolites. Nucleic Acids Res.
