PMML, the Predictive Model Markup Language, has been around for 18 years now, and has been adopted by many vendors of data mining software. The XML-based format is the de-facto standard for describing the results of the modeling phase, but also the preprocessing that is required before a prediction can be performed with the model.
KNIME is one of the leaders when it comes to processing PMML documents, making great efforts to always support the latest version of the format. PMML is baked right into KNIME, being the internal format that is used to transmit models between nodes. You may have noticed the blue connectors on model learners and data preprocessing nodes. These input and output ports are used to pass the PMML documents describing actions taken by the corresponding node. As one of few products, KNIME also supports PMML ensemble models, a feature that has been described in detail in another blog post. Reader and writer nodes for im- and exporting PMML complete the support and make it easy to share models with other software.
Learning a Model
Let's dive into KNIME for a small example of how PMML in KNIME works. In the following workflows we will use the iris data set, a commonly used set for data mining examples. The nodes we will use are in the packages KNIMELabs and Ensemble Learning.
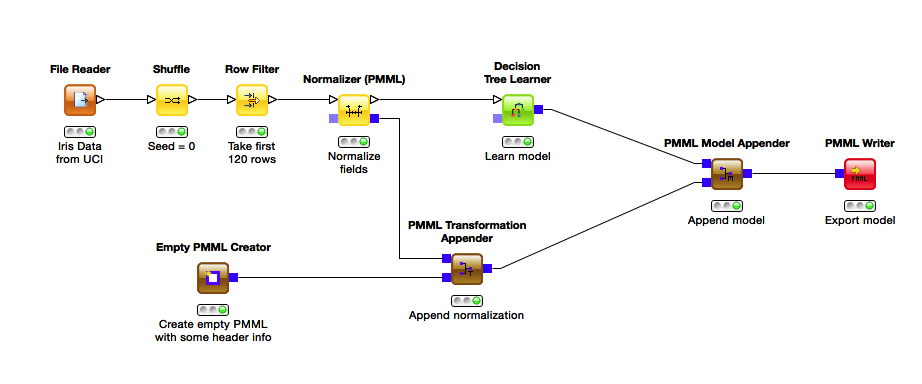
We first perform some preprocessing before feeding the data into a decision tree learner, which produces a decision tree model. Recently we made a change to how KNIME builds PMML documents, making it easier to put together the fragments that are produced by different nodes. You can see the result in the workflow above, where the data flows at the top and the corresponding PMML are built step by step at the bottom. This separation not only makes the workflow easier to understand, it also helps with building model ensembles.
The first step of PMML generation is the creation of an empty PMML document, without any preprocessing or model information. Here you can edit the document’s meta information, such as adding a copyright, description or custom annotations. To this document the transformations performed by the Normalizer are appended with a PMML Transformation Appender. This appender adds the information from the top input to the PMML document in the bottom input and performs checks to ensure the validity of the PMML, then passes it on to the next node. If you have multiple preprocessing steps, you can chain the appenders as often as you like. The final step of PMML generation is the PMML Model Appender, where the learned model is added to the document with the transformations. In our workflow we export the model to disk, but of course we could also pass it to one of the PMML predictor nodes to use the model directly on new data.
Learning an Ensemble of Models
The example becomes more complex when we want to build an ensemble of decision trees, where we need to learn many models on different data and then insert them all into a single document. Luckily KNIME has nodes to help you with that. The following workflow shows how this scenario is approached in KNIME.
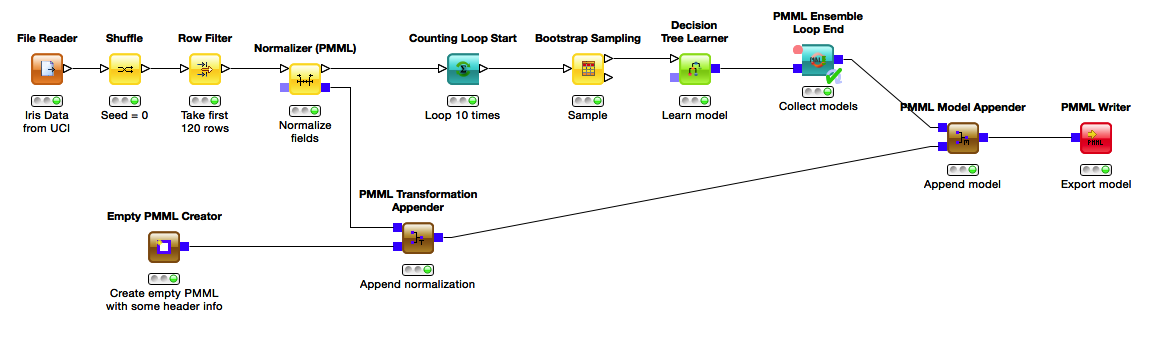
Here preprocessing and PMML initialization is the same as above, but the model generation is now much more complex: we loop 10 times, each time taking a bootstrap sample and learning a decision tree on it. The individual models are collected in a PMML Ensemble Loop End that outputs a PMML document containing all the models we have learned. Here we see how the separation of PMML creation and data processing helps us. Instead of passing the PMML into the loop, we append it to the ensemble model once it is finished. This avoids having the same preprocessing steps documented locally for each of the models in the ensemble. Instead, the preprocessing is appended once for all models. The workflow is not only easy to understand but also produces a smaller PMML document.
PMML Based Preprocessing and Prediction
Apart from producing PMML documents, KNIME is also a great choice for executing them on new data. There are multiple nodes for applying preprocessing and making predictions. The workflow below shows how simple it is to load a document, apply the preprocessing defined within and make a prediction.
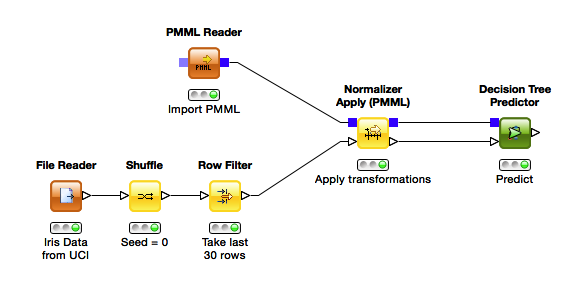
Despite its simplicity, there is a disadvantage to this approach: You need to know in advance which preprocessing methods were written to the PMML and which model type it contains. For the latter, there is the general PMML Predictor node, which takes any model type and calls the appropriate scoring code, but being able to look into the document to know which applier nodes to use might not be an option.
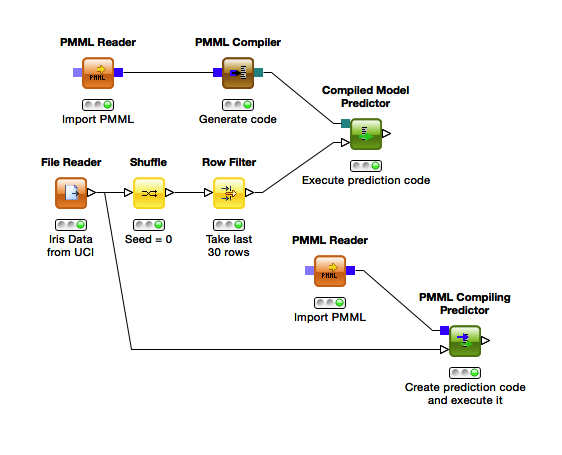
Rather new in KNIME and a good solution to this problem is the PMML Compiler, which was introduced in KNIME 2.11.2. It takes any PMML that was generated by KNIME and translates it into Java code. This code is passed to a special predictor, the Compiled Model Predictor, which executes it on new data. If you execute the model only once, you can also use the PMML Compiling Predictor. It combines the Compiler and the Predictor by compiling the PMML from its top input port and executing it directly on the data from the bottom input.
The compilation of PMML works well with models generated by KNIME, but also with PMML from a variety of other sources.
PMML Compilation
Apart from having preprocessing and prediction in a single node, the nodes that use PMML compilation are also faster than their traditional counterparts. You can, for example, expect an up to 4 times faster execution of decision tree models with the Compiled Model Predictor.
I hope this blog post has shed some more light on handling PMML documents in KNIME. PMML makes it very simple to share models among data mining software and is also a great format to store models for later execution. You can expect to hear more about the PMML compiler in the future, as we are planning to bring it to the cluster with a Spark PMML predictor. So stay tuned for more PMML news on knime.org.
The described example workflows can be downloaded from the attachment section of this article.
Requirements:
- KNIME Analytics Platform
- KNIME PMML Translation and Compilation extension (KNIME Labs)
- KNIME PMML Utility Nodes extension (KNIME Labs)
Further Reading:
