About a year ago we told a beautiful story about how KNIME Analytics Platform can be used to automate an established modeling process using the KNIME Model Factory. Recently our Life Science team faced an exhausting and frightening exercise of building, validating, and scoring models for more than 1500 data sets.
We want to share with you how we adapted the KNIME Model Factory for this monstrous application. We hope this will show you how to implement your own model building routines in the KNIME Model Factory. We will also demonstrate how to scale model building processes to very large tasks using KNIME Server Distributed Executors.
The Problem: Meet the Monster
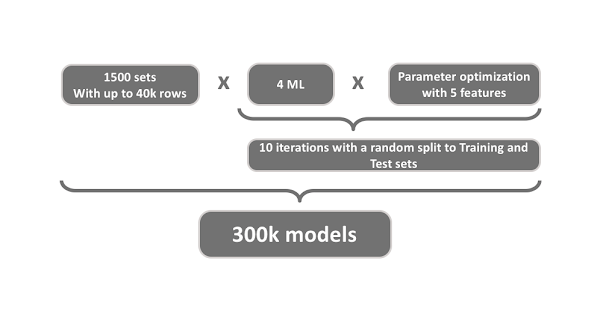
Our Life Science team wanted to apply their established modeling process to build and validate machine learning models for more than 1500 datasets. This modeling process is a set of workflows to extract the relevant data, transform it by eliminating artifacts and computing features, build models using different methods with tailored hyperparameter search, evaluate the models, select the best ones, and finally deploy them. We will further refer to these workflows as process workflows. For 1500 sets it would lead to more than 300 000 models (Figure 1)! That is much too large of a task to do manually, so we decided to use the KNIME Model Factory to help orchestrate the modeling process and scale it for this monstrous task.
Figure 1. The scale of the monstrous model building task: 1500 datasets, 4 ML models trained on each dataset, parameter optimization on 5 features on each model, 10 iterations for each optimization model, for a total of 300k models to train and evaluate.
Automatic Model Process Management: Meet the Beauty
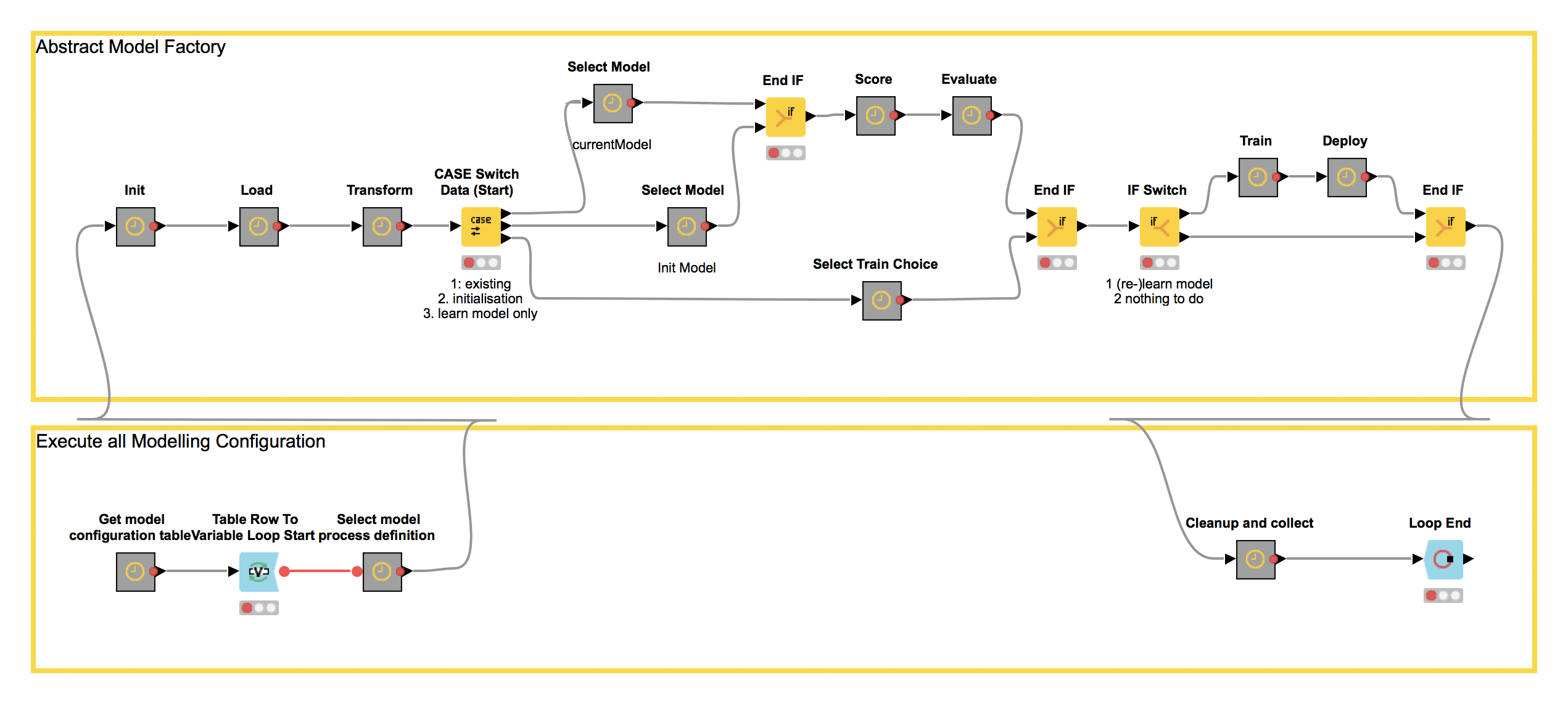
The KNIME Model Factory is a single application that orchestrates a set of user defined workflows for building models for diverse data sets (Figure 2). The flexible construction of the KNIME Model Factory allows you to set up different modeling processes for different datasets in a single application in a fully automated, reproducible, and easy to understand way. This is exactly what we, the Life Science team needed, so we adapted the Model Factory to implement and orchestrate their process workflows. You can find further detailed information on the setup of the abstract KNIME Model Factory in our white paper.
Figure 2. An workflow for the Model Factory to set up different modeling processes for different datasets in a single application in a fully automated, reproducible, and easy to understand way.
Creating the Monster Model Factory
Here is the final orchestration workflow, which we named the “Monster Model Factory” because of the monstrous size of the problem it’s designed to solve (and also because we liked the idea of working on a project with Monster in its name) (Figure 3). We will refer to this as the master workflow.
Figure 3. The master workflow for the Monster Model Factory to train and score a monster number of models.
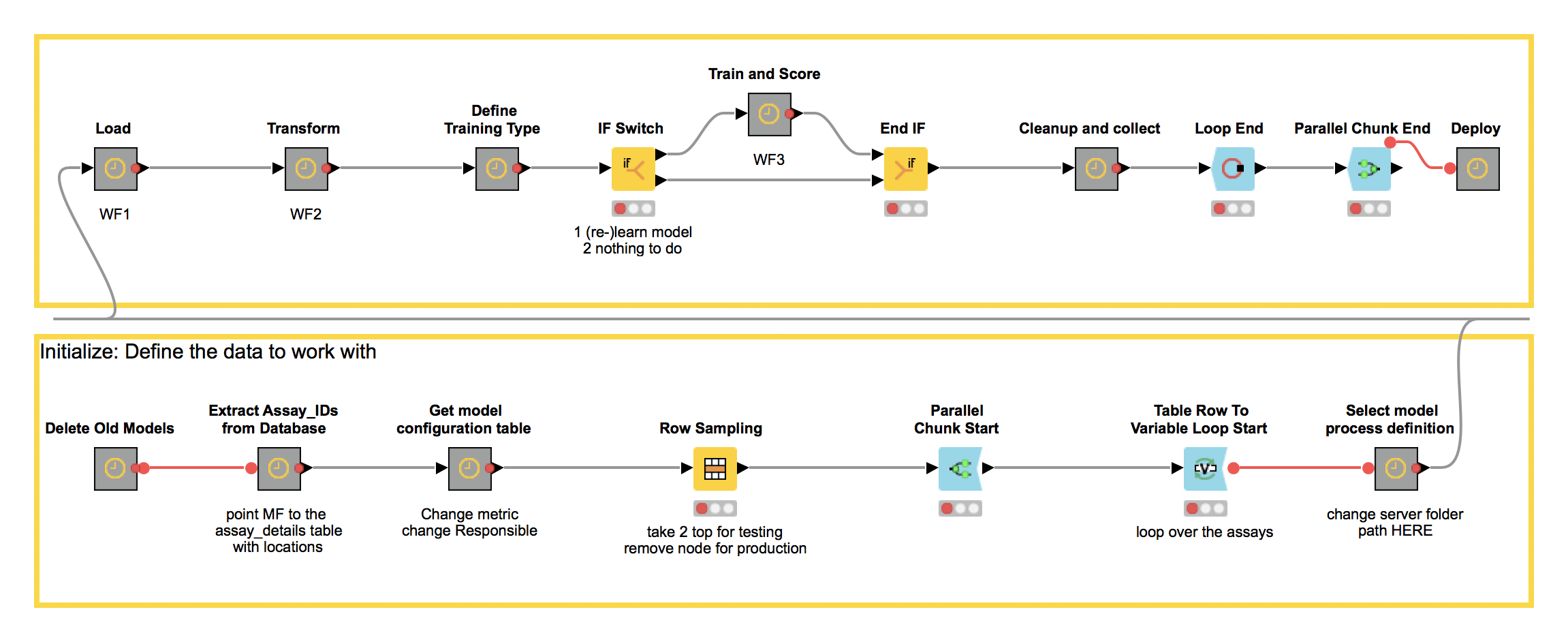
It starts in the lower part of the workflow, where the Model Management Process is initialized by specifying the metadata necessary to automate the modeling process. This metadata includes dataset ids, specific process workflows, the scoring criteria, and the URL of KNIME Server used for execution. In the upper part, the Monster Model Factory triggers the execution of the individual process workflows via a set of Call Remote Workflow nodes, which are encapsulated in the corresponding metanodes (Figure 4).
Our team incorporated three workflows as individual process steps in the Model Factory: Load, Transform, and Train and Score.
Figure 4. The contents of the Transform Metanode. It contains a Call Remote Workflow node, which triggers execution of the Transform workflow on KNIME Server.

Let’s have a look now at the three individual process workflows in detail.
1. The Load workflow extracts the data from a PostgreSQL database (Figure 5). Since the SQL queries are complex, this operation can take up to several hours for some of the datasets. This step could be skipped if the data are available locally (as in the workflow uploaded to KNIME Public EXAMPLES Server under 50_Applications/37_Monster_Model_Factory/Process/_Process_Step_Templates/workflows/Templates/01_Bioactivity_Prediction_Load_Local). Our team was working with ChEMBLdb, a large public repository of bioactivity datasets with information on molecules, proteins, and the relationships between them.
Figure 5. The process workflow 1: Load. Here data are extracted from a PostGreSQL database through a series of complex SQL operations.
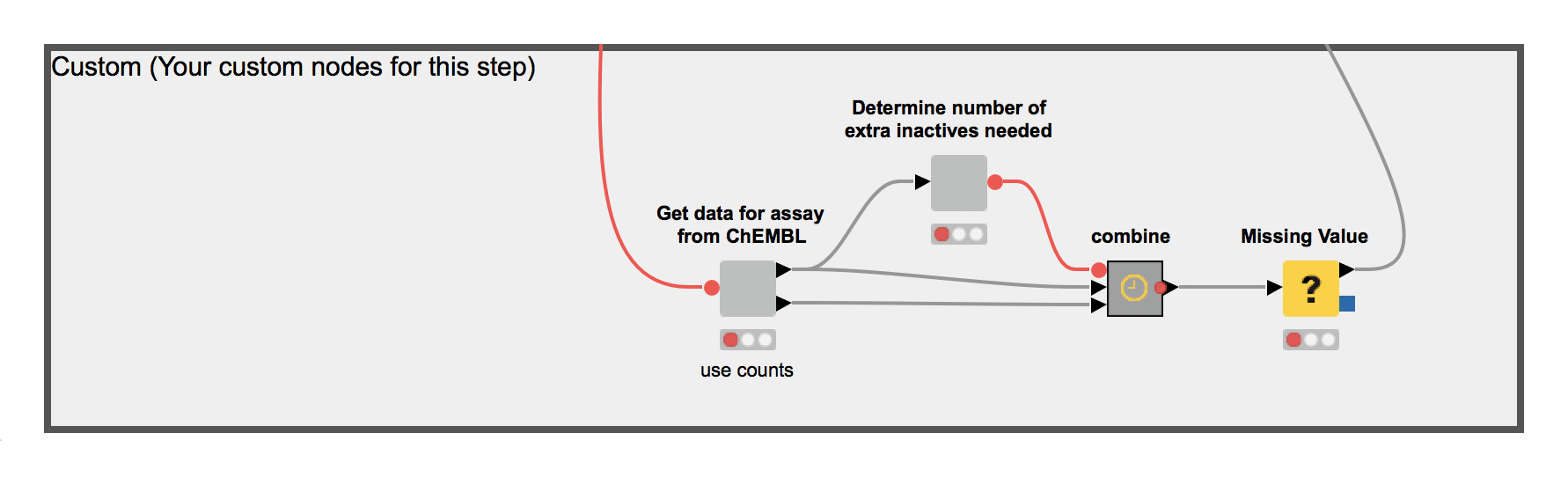
2. The Transform workflow uses the RDKit cheminformatics extension for KNIME to preprocess chemical structures and compute chemical fingerprints, i.e. create a machine readable representation of the molecule from the encoded structure in the dataset, which will be used to train a machine learning model (Figure 6).
Figure 6. The process workflow 2: Transform. This workflow creates a machine readable representation of the molecule from the encoded structure in the data set.
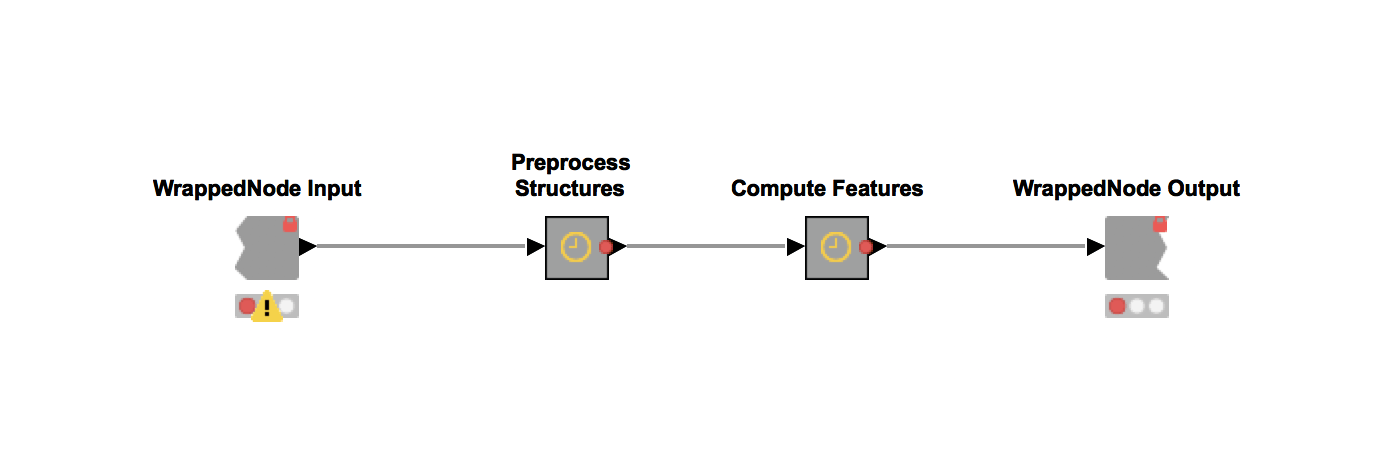
3. The Train and Score workflow is the most computationally expensive part of the Model Factory (Figure 7). This workflow trains four models with four different machine learning algorithms and optimizes all four of them with a parameter optimization procedure. It is based on a workflow from our recent blogpost on Parameter Optimization. This training step is repeated ten times. Next, the forty trained models (4 ML algorithms x 10 iterations) are scored using the test data and the model with the highest performance is picked for deployment.
Figure 7. The process workflow 3: Train and Score. This workflow trains four models with four different machine learning algorithms and optimizes all four of them with a parameter optimization procedure. The training is repeated ten times. The model with the highest performance is picked for deployment.

The best models and their statistics, along with the statistics of all models that were generated in the Train workflow for a dataset, are saved in the Deploy Metanode of the Monster Model Factory master workflow.
Scaling the Monster
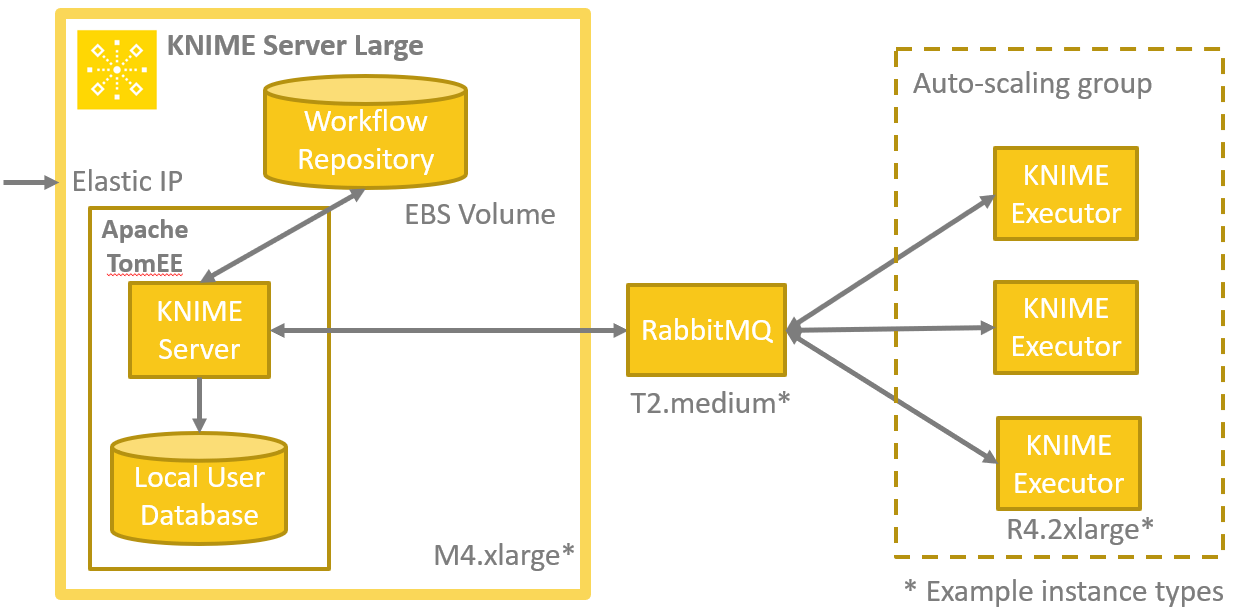
Now that we’ve shared with you how we automated the modeling process for a single dataset, let’s explore how we can scale it up to our Monster Project with 1500 sets. Running that computation on a single machine just isn’t practical, it would take too long. That’s where the new Distributed Executors feature of KNIME Server comes into play. Using a template to deploy the whole infrastructure in the cloud using Amazon Web Services (AWS) takes just a couple of minutes, and allows adding more executors to the calculation as needed using an auto-scaling group. (Figure 8).
Figure 8. Deployment of the Monster Model Factory on Amazon Web Services (AWS)
To take advantage of the multiple available Distributed Executors, we added a Parallel Chunk Loop around the process workflows in the master workflow of the Monster Model Factory (Figure 3). This triggers execution of multiple jobs in parallel on separate executors . To make sure the monster didn’t burn up the cluster, we defined the maximum size of chunk in the Parallel Chunk Start node to be the maximum number of executors we wanted to have running at a time.
Executing the Monster Model Factory
You start the execution of the Monster Model Factory by launching the job for the master workflow on KNIME Server. In a few seconds it triggers execution of the process workflows. Depending on the number of available executors, you will see several jobs starting and running in parallel on the server (Figure 9). Once a job is finished, the Factory starts the next job in the queue.
Figure 9. Master and process workflows executing on KNIME Server. Notice the data and workflow structure in the KNIME Explorer panel of the server. Also notice the workflow jobs running in parallel in the workflow section during execution of the Monster Model Factory.
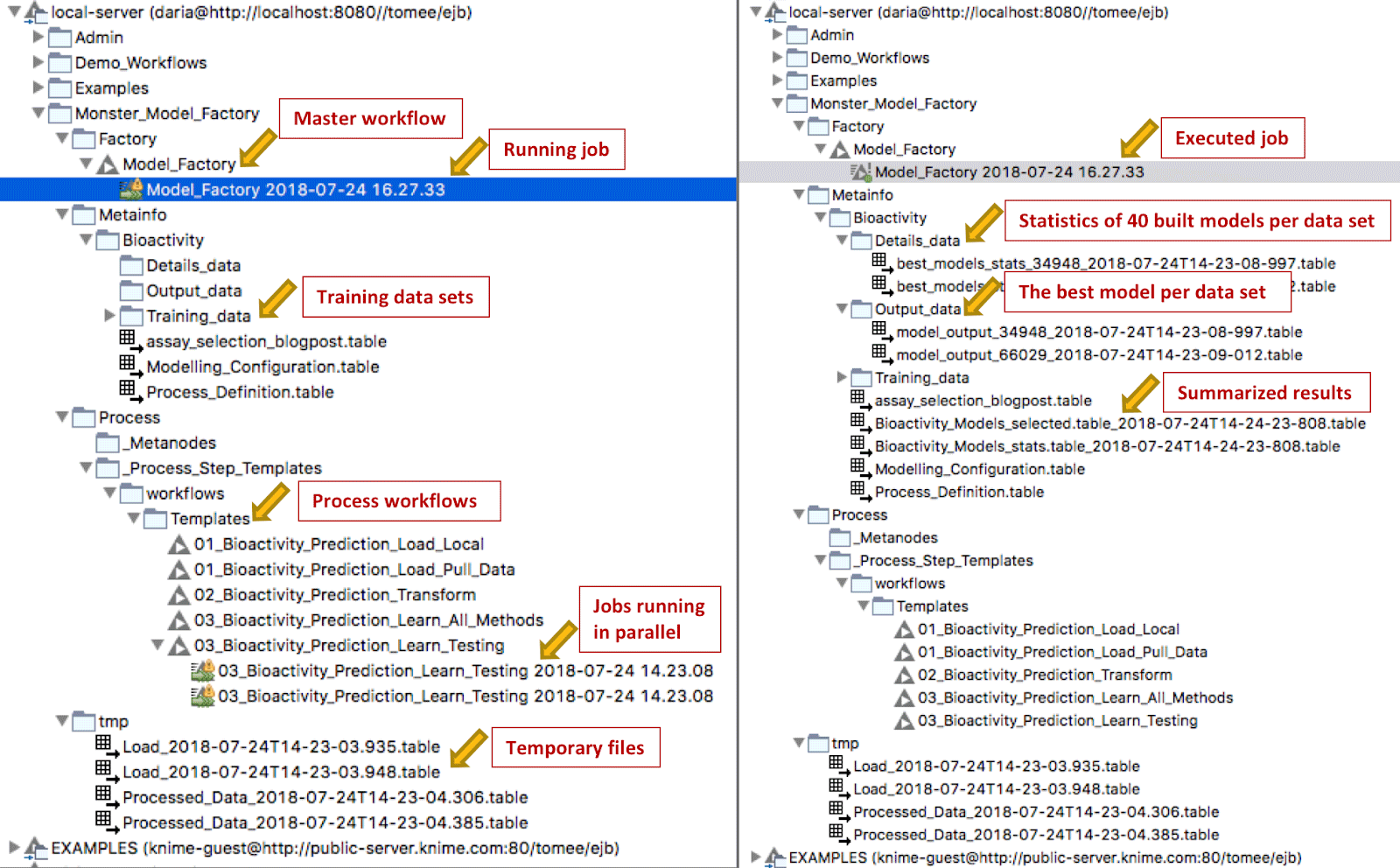
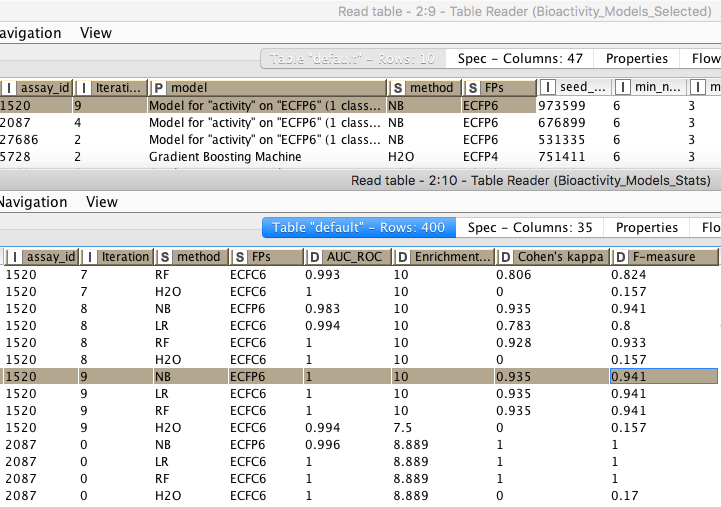
During execution of the factory, the temporary files it creates are stored in the tmp folder. These are removed once a new instance of the Factory starts. We can find results for each dataset in the Metainfo directory. In the Details folder we find statistics for all the models generated during the training step; the Output folder contains the final best model for each dataset. Once the Factory has finished executing the final results are summarized in two tables: Bioactivity_Models_Selected and Bioactivity_Models_Stats (Figure 10).
Figure 10. Overview of Bioactivity_Models_Selected and Bioactivity_Models_Stats data tables containing the final results.
To sum up, we have shared with you how to automate the model building process with KNIME Analytics Platform and scale it to build thousands of models in a reasonable amount of time using KNIME Server Distributed Executors. If you would like to use the Monster Model Factory locally, just exchange the Call Remote Workflow node with the Call Local Workflow node and enjoy the automation.
We hope you find the Monster as useful as we do!
- Download the Monster Model Factory workflows from the KNIME Hub here.
- The Video is available on YouTube via https://www.youtube.com/watch?v=PLfkR7RHbeY
