In this blog series we’ll be experimenting with the most interesting blends of data and tools. Whether it’s mixing traditional sources with modern data lakes, open-source devops on the cloud with protected internal legacy tools, SQL with noSQL, web-wisdom-of-the-crowd with in-house handwritten notes, or IoT sensor data with idle chatting, we’re curious to find out: will they blend? Want to find out what happens when IBM Watson meets Google News, Hadoop Hive meets Excel, R meets Python, or MS Word meets MongoDB?
Today: SAS, SPSS, and MATLAB meet Amazon S3: setting the past free
I am an old guy. And old guys grew up with proprietary data formats for doing cool data science things. That means I have literally 100s of SAS, SPSS and MATLAB files on my hard disk and I would still love to use that data - but I no longer have the cash for the yearly licenses of those venerable old packages.
I know I can read all those files for free with KNIME. But what I REALLY want to do is move them into an open format and blend them somewhere modern like Amazon S3 or Microsoft Azure Blob Storage. But when I check out the various forums, like the SAS one, I find only horrendously complicated methods that - oh by the way - still requires an expensive license of the appropriate legacy tool. So it’s KNIME to the rescue and to make it easy, this example pulls files of all three legacy types and allows you to either move them or first convert them to an open source format before moving them to - in this example - Amazon S3.
It’s easy, it’s open, it’s beautiful…. and it’s free.
Topic. Recycling old data files from legacy tools.
Challenge. Transfer data from SAS, SPSS, and Matlab into Amazon S3 data lake passing through KNIME.
Access Mode. Access nodes to SAS, SPSS, and Matlab files and connection node to Amazon S3.
The Experiment
First, you will need your credentials or Access Key/Secret Key for Amazon S3. If you don’t have one or are new to Amazon S3, the folks have made it easy for you to sign up and try for free. Just follow the instructions here: www.amazon.com/S3.
By the way, to be totally inclusive, KNIME also supports Microsoft Azure Blob Storage, and they also offer a free trial account. So sign up for both and compare them. For purposes of keeping this blog short, I will restrict myself to S3.
R is used to read the SPSS and MATLAB files. If you do not have the KNIME R extensions, you will want to install them via the File/Install pulldown within KNIME. By the way, the MATLAB package is not automatically included, but the workflow takes care of this one-time installation for you.
The Workflow
An overall view of the workflow is here:
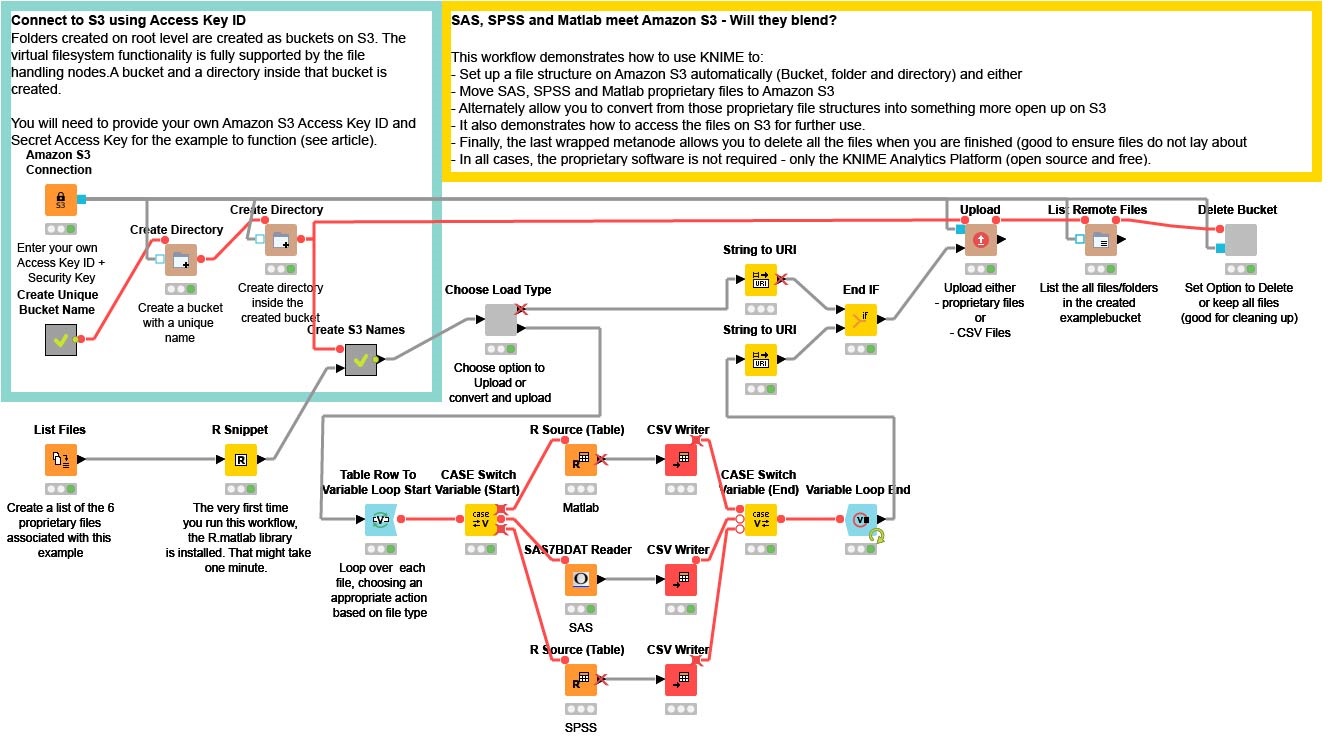
In the upper left hand corner, I first enter my Amazon S3 credentials into the Amazon S3 Connection node. This establishes a valid connection to S3. By default, there is just “space” on S3 and I need to define what is known as a Bucket and then establish a Directory within that Bucket for all my files. I do that in the upper left hand corner by randomly generating a name for the Bucket. This is helpful as I may do a lot of testing and playing with this workflow, and I want to see the various flavors of files that I produce. For a production application, this section would be replaced with appropriate established String values.
I then want to find the SAS (.SASB7DAT), SPSS (.SAV) and Matlab (.MAT) files. In this example workflow, I’ve provided a directory with a small variety of each file type actually contained within the workflow location. That List Files node creates that list for me. In preparation for uploading, I need to convert local PC filenames into more generic URI names (which are recognized by S3) and that is what I do in the small metanode named “Create S3 Names”.
I then have a choice to make. I can either move the proprietary files directly onto S3, or I can first convert each of those files into an open format before moving them to S3. I do that with the small wrapped metanode named “Choose Load Type”. Simply by right clicking that node, I get a specific configuration box (made with Quickforms) that allows me to choose.
If I choose “Proprietary”, then the top branch is chosen, the appropriate files name are used and then KNIME does a bulk-load of the files into my Amazon S3 directory. Extremely easy and efficient but, at the end of the day, still in that proprietary form.
If I choose “Open (CSV Format)”, I first read each of the proprietary files with the appropriate KNIME node. In this case, we have a native node for SAS7BDAT files and two R nodes that use the appropriate R package to read SPSS and MATLAB files. I then use CSV Writer nodes to write each file. Note the LOOP and CASE SWITCH construct, which takes each row and - based on the file type - automatically uses the appropriate READ node.
When the conversion is finished, all those now Open Format CSV files are bulk loaded into S3. At the end of the flow, I do a simple LIST Directory to show which files are there.
As a very last step, I have a small wrapped metanode that allows me to choose whether to delete the Bucket I created - very handy when cleaning up after my experiments.
Note. If you end your KNIME session and want to come back to this example, you will each time need to reset the workflow then rerun (so that you get a NEW S3 Bucket and Directory each time).
The Results
Will they blend? Absolutely. In the figure below, you can clearly see the uploaded .csv files in the list of remote files from the cloud:
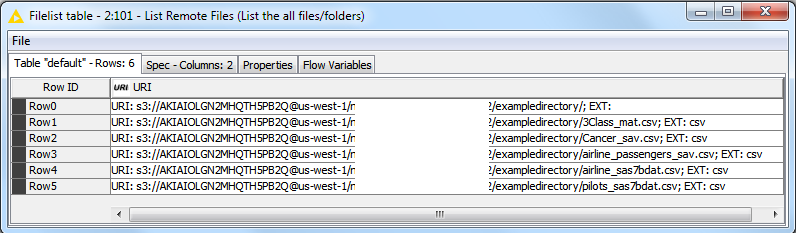
It is actually fun to play with this example KNIME workflow. You can easily point to your own files or even change the File Readers to your favorite other type of data source (proprietary and old or otherwise). What I find extremely powerful about this blending workflow: you can also change the output data type. Do you fancy NoSQL, Redshift, XML, Excel or any other format? Simply change to the appropriate Writer (either native, ODBC based or R based). And if you want to check out Microsoft AZURE, you can easily modify this workflow by changing the Amazon S3 Connection node to an Azure Blob Store Connection node, with the appropriate parameters, and the workflow should work on Azure as well.
Although these sample files are small, this workflow will work for extremely large files as well since - after all - that is what the cloud is for.
Indeed, in KNIME Analytics Platform you can create streamed wrapped metanodes that subset and manipulate your data “on the fly” creating fantastic opportunities to efficiently use KNIME on those ever growing Data Lakes within the cloud.
Note. If you use a supported streaming data types, such as CSV or KNIME Table, you do not need to bulk-download the S3 file but can transfer the data via streaming. For examples, search the KNIME Examples server with the keyword S3 or Azure.
This workflow is available, without the API keys (of course), on the KNIME Hub from the 01_Data_Access folder.
Coming Next…
If you enjoyed this, please share this generously and let us know your ideas for future blends.
We’re looking forward to the next challenge. Next time we will attempt a data blending in English literature. We will attempt to blend Kindle epub documents and JPEG images while trying to reunite the Montague and Capulet families. Which one will be harder?
