In this blog series we’ll be experimenting with the most interesting blends of data and tools. Whether it’s mixing traditional sources with modern data lakes, open-source devops on the cloud with protected internal legacy tools, SQL with noSQL, web-wisdom-of-the-crowd with in-house handwritten notes, or IoT sensor data with idle chatting, we’re curious to find out: will they blend? Want to find out what happens when IBM Watson meets Google News, Hadoop Hive meets Excel, R meets Python, or MS Word meets MongoDB?
Today: OCR on Xerox Copies meets Semantic Web. Have Evolutionary Theories changed?

The Challenge
Scientific theories are not static over time. As more research studies are completed, new concepts are introduced, new terms are created and new techniques are invented. This is of course also true for evolutionary theories. That is, evolutionary theories themselves have evolved over time!
In today’s challenge we are going to show how the theory of evolution has evolved from the first Darwin’s formulation to the most recent discoveries.
The foundation stone of evolutionary biology is considered to be the book “On the Origin of Species” (1859) by Charles Darwin. This book contains the first revolutionary formulation of the theory of evolutionary biology. Even though the book at the time has produced a revolution in the approach to species evolution, many of the concepts illustrated there might seem now incomplete or even obsolete. Notice that it was published in 1859, when nothing was known about DNA and very little about genetics.
In the early 20th century, indeed, the Modern Synthesis theory reconciled some aspects of Darwin’s theory with more recent research findings on evolution.
The goal of this blog post is to represent the original theory of evolution as well as the Modern Synthesis theory by means of their main keywords. Changes in the used keywords will reflect changes in the presented theory.
Scanned Xerox copies of Darwin’s book abound on the web, like for example at http://darwin-online.org.uk/converted/pdf/1861_OriginNY_F382.pdf. How can we make the contents of such copies available to KNIME? This is where Optical Character Recognition (OCR) comes into play.
On the other side, to find a summary of the current evolutionary concepts we could just query Wikipedia, or better DBPedia, using semantic web SPARQL queries.
Xerox copies on one side, read via OCR, and semantic web queries on the other side. Will they blend?
Topic. Changes in the theory of evolution.
Challenge. Blend a Xerox copy of a book with semantic web queries.
Access Mode. Image reading, OCR library, SPARQL queries.
The Experiment
Reading the Content of a PNG Xerox Copy via Optical Character Recognition (OCR)
Darwin’s book is only available in printed form. The first part of our workflow will try to extract the content of the book from its scanned copy. This is only possible using an Optical Character Recognition software.
We are in luck! KNIME Analytics Platform integrates the Tesseract OCR software as the KNIME Image Processing - Tess4J Integration. This package is available under KNIME Community Contributions - Image Processing and Analysis extension (see how to install KNIME Extensions). Let’s continue step by step, after the package installation.
- The Image Reader node reads the locally stored image files of the pages of the book “On the Origin of Species”.
- The read images are sent to the Tess4j node, which runs the Tesseract OCR library and outputs the recognized texts as Strings, one text String for each processed PNG page file.
- Each page text is then joined with the corresponding page image, converted from ImgPlusValue to PNG format in a KNIME PNGImageCell data cell. The goal of this step is to allow later on for visual inspection of the processed pages via the Image Viewer node.
- Notice that only the “Recap” and the “Conclusions” sections of Darwin’s book are processed. That should indeed be enough to extract a reasonable number of representative keywords.
This part of the workflow is shown in the upper branch of the final workflow in figure 1, the part labelled as “Optical Character Recognition of Xerox Copy”.
Figure 1. Final workflow where OCR (upper branch) meets Semantic Web Queries (lower branch) to show the change in keywords of original and later formulations of the theory of evolution. This workflow is available on the KNIME EXAMPLES server under 99_Community/01_Image_Processing/02_Integrations/03_Tess4J/02_OCR_meets_SemanticWeb99_Community/01_Image_Processing/02_Integrations/03_Tess4J/02_OCR_meets_SemanticWeb*
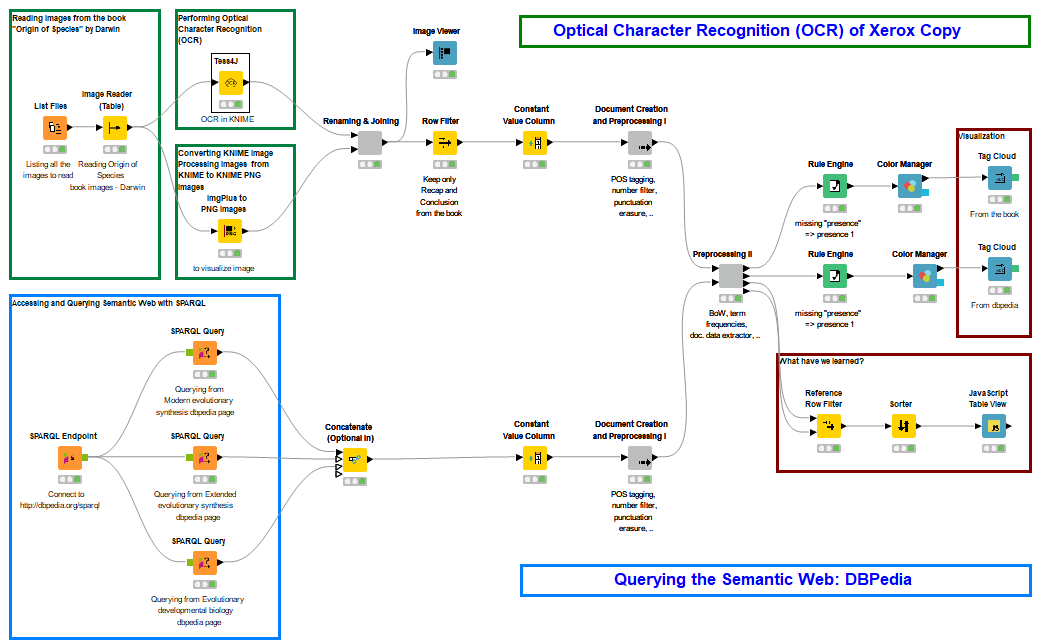
(click on the image to see it in full size)
Querying the Semantic Web: DBPedia
On the other side, we want to query DBPedia for the descriptions of the modern evolutionary theories. This part can be seen in the lower branch of the workflow in figure 1, the one named “Querying the Semantic Web: DBPedia”.
- First, we establish a connection to DBpedia SPARQL endpoint: http://dbpedia.org/sparql.
- Then we make three queries on the 3 pages “Modern evolutionary synthesis”, “Extended evolutionary synthesis” and “Evolutionary developmental biology” respectively.
- The results from the 3 queries are collected with a Concatenate node.
Blending Keywords from the Xerox Copy and from the Semantic Web Queries
We have now texts from the book and texts from DBPedia. We want to distill all of them to just a few representative keywords and blend them together.
- Both the two branches of the workflows pass through “Document Creation and Preprocessing I” wrapped metanode. Here standard text processing functions are applied, such as: case conversion, punctuation removal, number filtering and POS tagging.
- In the following wrapped metanode, named “Preprocessing II”, the terms are extracted and their relative frequencies are computed. The two lists of terms are then joined together. The column “presence” marks the terms common to both datasets with a 2 and the terms found in only one dataset with a 1.
- The two tag clouds are created, one from the terms in Darwin’s book and the other from the terms in the DBpedia search results. Words common to both datasets are colored in red.
- Finally, we can isolate those innovative terms, used in the description of the new evolutionary theories but not in Darwin’s original theory. This is done with a “Reference Row Filter” node and displayed with a “Javascript Table View”.
The final workflow is available on the EXAMPLES server under:
99_Community/01_Image_Processing/02_Integrations/03_Tess4J/02_OCR_meets_SemanticWeb99_Community/01_Image_Processing/02_Integrations/03_Tess4J/02_OCR_meets_SemanticWeb*
The Results
Figure 2 shows the tag cloud with the terms from Darwin’s book “On the Origin of the Species”, while figure 3 shows the tag cloud from the terms found in the results from DBPedia queries.
Natural Selection is a central concept in Darwin’s evolutionary theory and this is confirmed by the central position of the two words, “natural” and “selection”, in the tag cloud in figure 2. The two words are in red. This means that the same terms are also found in modern evolutionary theories, even though in a less central position inside the tag cloud (Fig. 3).
Interestingly enough the word “evolution” is not found in Darwin’s book. Although this term was soon associated with Darwin’s theories and became popular, Darwin himself preferred to use the concept of “descent with modification” and even more “natural selection”, as we have remarked earlier.
Words like “species” and “varieties” also play a central role into Darwin’s theory. Indeed, the whole theory spawned from the observation of the species variety on Earth. On the opposite words like “modern”, “synthesis”, and “evolution” are the cornerstone of the modern evolutionary theories.
Figure 2. Word Cloud generated from the “Recap” and “Conclusions” sections in the book “On the Origin of the Species” by Charles Darwin

(click on the image to see it in full size)
Figure 3. Word Cloud generated from results of DBPedia queries: “Modern evolutionary synthesis”, “Extended evolutionary synthesis” , and “Evolutionary developmental biology”

(click on the image to see it in full size)
What has been learned from the publication time of the book “On the Origin of Species” to the current time? One thing that Darwin for sure could not know is genetics!
If we look for the word “gene” in the table view of the Javascript Table View node, we surely find “genetics”, “gene”, and a number of other related words! Remember that this table view displays the terms found in the description of modern evolutionary theories but not in Darwin’s original book (Fig. 4).
Figure 4. List of “genetics” related words present in the modern evolutionary theories (as derived from the queries to DBPedia) and not present yet in Darwin’s original book (as derived from the OCR processing of the scanned PNG images of the book pages). Such words are all listed in the table view of the Javascript Table View node in the final workflow.
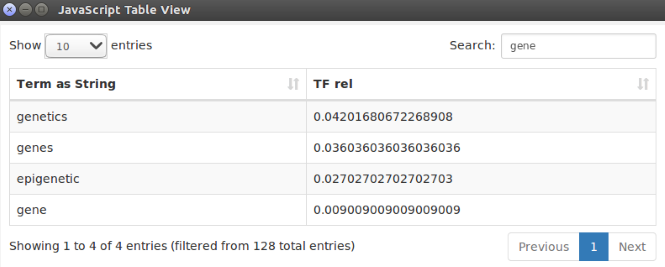
From the results of this small experiment we understand that the evolutionary theory has itself evolved from Darwin seminal work to the modern research studies.
Also, in this experiment, we successfully blend data from a scanned Xerox copy and data from DBPedia. We have used the Tesseract OCR integration to extract the book content and SPARQL queries to extract DBPedia descriptions … and yes, they blend!
Coming Next …
If you enjoyed this, please share it generously and let us know your ideas for future blends.
We’re looking forward to the next challenge. For the next challenge we will tackle the Spreadsheet world, by trying to blend an Excel Sheet with a Google Sheet. Will they blend?
* The link will open the workflow directly in KNIME Analytics Platform (requirements: Windows; KNIME Analytics Platform must be installed with the Installer version 3.2.0 or higher)
