Keeping track of the latest developments in research is becoming increasingly difficult with all the information published on the Internet. This is why Information Extraction (IE) tasks are gaining popularity in many different domains. Reading literature and retrieving information is extremely exhausting, so why not automate it? At least a bit. Using text processing approaches to retrieve information about drugs has been an important task over the last few years and is getting more and more important1.
In a previous blog post Fun with Tags, we looked at how to how to train a named-entity recognition model to detect diseases in biomedical literature with KNIME Analytics Platform. This time, instead of disease names, we want to create a model that automatically detects drug names.
In addition, we will go one step further and predict the purpose of those drugs detected by the trained model. This can help to get an understanding of the drugs mentioned in articles of interest and might also help in studies about drug repurposing. Has this drug lately been mentioned together with other drugs, although they have little in common? Could there be an unknown or new connection which might help to use the drug for another purpose than usual?
Specifically, we will train a named-entity recognition (NER) model to detect drug names in biomedical literature. To do this we need a set of documents and an initial list of drug names. Since our goal is not only the recognition of these drug names but also the prediction of a drug’s purpose, we need some additional information about these drugs.
After collecting the list of drug names, we will automatically extract abstracts from PubMed. These documents will be split in two parts: one part used as our training corpus to train the model and one part for testing and validation purposes. The final model is then used to tag the whole set of documents. Based on the tagged drug names, we will create a drug-drug co-occurrence network. All of our known drugs (the drugs from our initial list) will have some additional information which can be used to predict the purpose of a drug that recognized for the first time by our model (and was not in our initial list).
The work was split into four different workflows.
- Gathering drug names and related articles
- Preprocessing, Model Training and Evaluation
- Create a Co-Occurrence Network and Predict Drug Purposes
- Extract Interesting Subgraphs
Gathering drug names and related articles
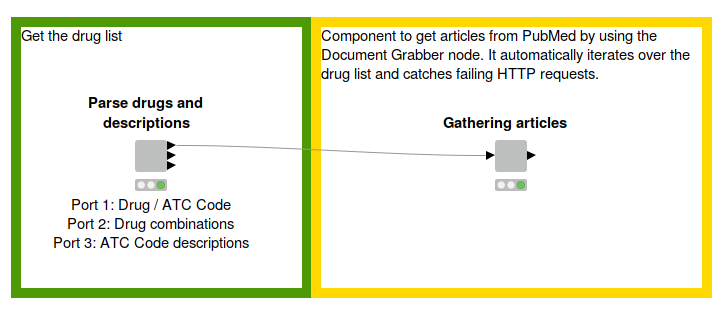
Fig. 1: Workflow to parse the drug list and descriptions from the WHO website and create a corpus by using the Document Grabber to fetch articles from PubMed. The functionality is wrapped in components for better clarity. Download the Creating a Corpus workflow from the KNIME Hub here.
Dictionary creation (Drug names)
As mentioned above, our initial list of drug names should have some sort of additional information which can be used to predict the purpose of newly identified drugs. Therefore, we decided to use drugs that are covered by the Anatomical-Therapeutic-Chemical (ATC) Classification System2, which is published by the World Health Organization. It contains around 800 drug and drug combinations whereas each drug is associated to one or more ATC codes.
ATC Classification System
The ATC code itself consists of seven letters and is separated into five different levels. As an example there is acetylsalicylic acid (aspirin) with ATC codes A01AD05, B01AC06 and N02BA01. The first letter stands for one of fourteen anatomical main groups which will be used for ATC code prediction later. The succeeding two letters describe the therapeutic subgroup, followed by one letter describing the therapeutic/pharmacological subgroup. The fourth level is resembled by the fifth letter and stands for the chemical/therapeutic/pharmacological subgroup.
The last two digits then indicate the generic drug name. For example A01AD05, there is A for alimentary tract and metabolism, A01 for stomatological preparations, A01AD for other agents for local oral treatment and finally A01AD05 for the compound’s name acetylsalicylic acid. More information about the different ATC levels and meanings can be found here (https://en.wikipedia.org/wiki/Anatomical_Therapeutic_Chemical_Classification_System).
Corpus creation
In the next step, we can start to gather abstracts related to the drugs from our drug list. As a source for biomedical literature we chose the widely-known PubMed database. To retrieve articles from PubMed and put them into KNIME we can use the Document Grabber node. It fetches a certain number of articles for each provided drug name in our drug list. In this case we try to get 100 articles per drug name.
Preprocessing, model training and evaluation
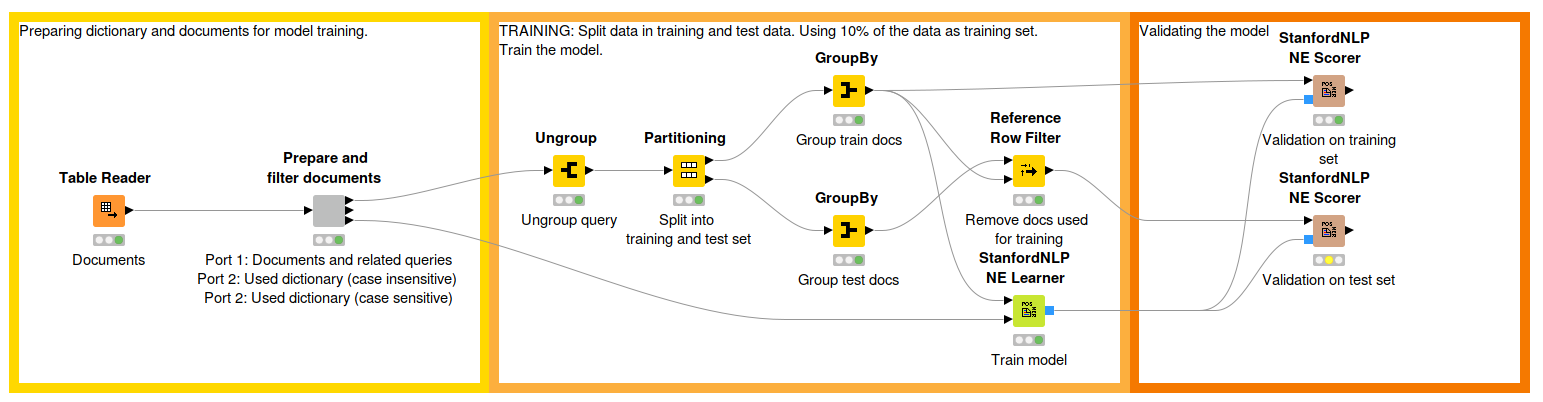
Fig. 2: Workflow that describes the model training process. The first part reads the text corpus created in the first workflow and preprocesses and filters some articles. The middle part shows the model training while the third part is for evaluation. Download the workflow, Traine a NER Model, from the KNIME Hub here.
To guarantee the quality of our corpus, some preprocessing tasks are required.
At first we check, if the downloaded articles contain the query term. PubMed sometimes provides articles that are related to a drug with a similar name, but does not exactly contain the search query. Since we can’t prepare the abstracts for model training then, we filter the unrelated articles. This can be done by tagging the articles with the Wildcard Tagger node and removing an article in case no word could be tagged. Afterwards we remove all drugs from our drug list with less than 20 remaining articles to ensure a dataset with enough sample sentences containing the drug names. This yielded a final text corpus of 44891 unique abstracts (53311 with duplicates) with 207875 annotated drug named-entities in total.
Now, we can start to train our model, but before we do this, we partition our data into a training and a test set. We used 10% of the data for the training set (approx. 5000 articles). To train the model we use the StanfordNLP NE Learner node which currently provides 15 different parameters. Some important parameters are the useWord (set to true), useNGrams (set to true), maxNGramLength (set to 10) and noMidNGrams (set to true) parameters. They define whether to use the whole word, as well as substrings of the word as a feature, how long these substrings can be and whether the substrings can only be used as feature if they are taken from the beginning or end of the word. These things might not always be relevant, but in terms of drug names which often share similar word stems, it’s quite useful.
Another important setting is the case sensitivity option of the learner node. Since we don’t know which case is used for the words within our corpus, we choose case insensitivity, so that no matter which case is used, it is labeled by the learner node.
After training the model, we can evaluate the model by using the StanfordNLP NE Scorer. It tags the documents once by using regular expressions and once by using the trained model and compares the tags. The resulting table provides basic scores like precision, recall and counts for true and false positives/negatives.
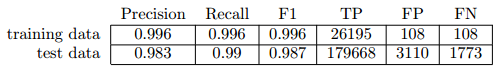
Table 1: This table shows the number of true/false positives, false negatives as well as some basic metrics like precision, recall and f1.
As we can see the majority of drug names could be tagged correctly. False positives are not necessarily a problem, because the model is not only for tagging known drug names from our initial drug list, but it also generalizes to find new entities. Otherwise we could have just used the Wildcard or Dictionary Tagger. However, regarding false positives, we still don’t know if the newly tagged words, are drug names at all, but we will investigate it later.
Since the Scorer node only gives us counts and scores, we use an additional approach for evaluation which helps us to identify the words causing false positives and negatives. Basically, we do what the StanfordNLP NE Scorer node does internally. We tag the documents twice, once using the StanfordNLP NE Tagger and once using the Wildcard Tagger. Afterwards we count the number of annotations for each drug and compare the different tagging approaches. For most drugs, we can see that they were annotated at the same frequency, no matter which annotation method was used. For the example of insulin, we can see that the model sometimes just tagged insulin although there was another name component (e.g. aspart or degludec).
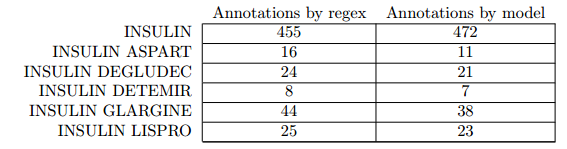

Table 2: This table shows the number of annotations for each insulin related drug by using regex and the trained model . As we can see, the model annotated insulin more often than actually available in the literature since it failed to detect the second part. This helps to identify the amount of false positives from Table 1.
Apart from all the measurements, we of course want to know what kind of newly identified entities there are. To get a small overview, we can use the String Matcher node to identify similarities between new words and drug names from our initial list. After doing so, we see that there are some words that are just spelling mistakes or slight variations of the drug name due to to different spellings in other countries. Some newly found names were just extensions of known drugs (e.g. insulin isophane). However, in the end we were able to detect around 750 new words whereas more than half could not be linked to a drug name from the initial list. These words would need further investigation.
Create a co-occurrence network and predict drug purposes
Enough about training and evaluating the model. Let’s make use of the model. We can use the drug names tagged by our model to create a co-occurrence network of drug names co-occuring in the same documents. This allows us to investigate the newly found drug names in more detail and, furthermore, enables the prediction of the purpose of those newly identified drugs.. To create that network, we use the Term Co-occurrence Counter node which counts co-occurrences on sentence or document level. In this case it is enough to set it to document level, since our documents are high-level abstracts and it’s very likely that drugs being named together in an abstract are somehow related. Based on our resulting term co-occurrence table, we can create a network.
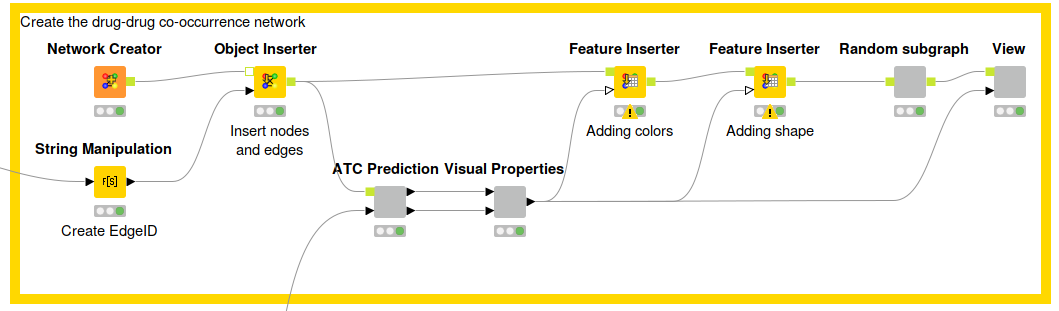
Fig. 5.: Workflow that describes the network creation process. First, we use the Network Creator node to create an empty network that can be filled with new nodes and edges by using the Object Inserter. Afterwards, we predict the ATC codes and create visual properties (color & shape) for the nodes within the network. These properties can be added by using the Feature Inserter node. After doing so, we use the Network Viewer JS (hidden in the View component) to visualize the network. Download the Co-occurence Network workflow from the KNIME Hub here.
This network can now be used to predict ATC codes of drugs. For each of our newly identified drug names, we do a majority vote, meaning that we set the ATC code that occurred most frequently in the neighborhood of an “unknown” drug. For visualisation purposes, all drugs in the network are colored based on the first level of the ATC code. Additionally, newly detected drugs are displayed as squares and known drugs as circles, respectively. This helps to evaluate and comprehend the prediction of the ATC code. As mentioned before, our initial list had around 800 drug names and the list of newly found entities contains 750 drugs. So in total there are quite some nodes in the network which makes the view pretty confusing. To avoid this, I show you how to extract relevant subgraphs to evaluate and comprehend predictions in the next section.
Extract interesting subgraphs
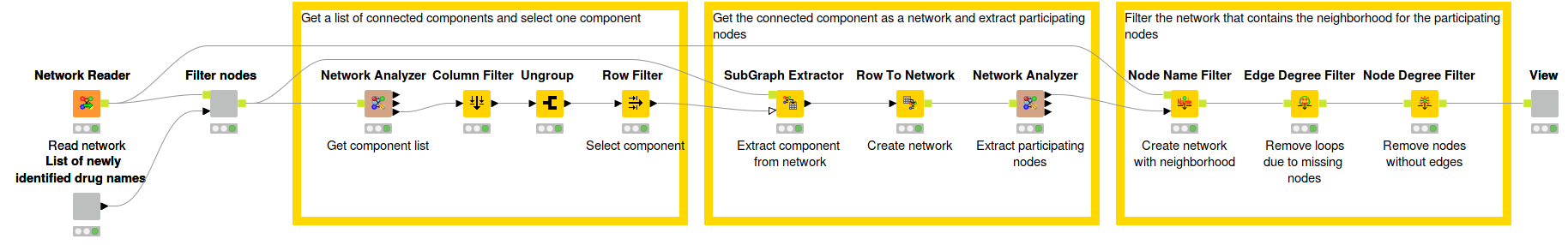
Fig. 6: Workflow that describes the process of extracting small connected components to investigate the predictions.
To investigate the predictions in detail, we can use connected components of newly detected drugs. At first, we remove all drugs from the network that are in the initial drug list to get a set of nodes in the network containing only the newly identified drugs. Afterwards, we re-add all of the previously filtered drugs that are in the first neighborhood of drugs from the component we are looking at. In the end, each connected component consists of a set of co-occurring newly detected drug names plus their neighbors from the initial network. This approach makes evaluating easier since we first filter mostly drugs from the initial drug list that tend to be connected to a huge number of drugs, but later re-add these to a smaller set of unknown drugs.
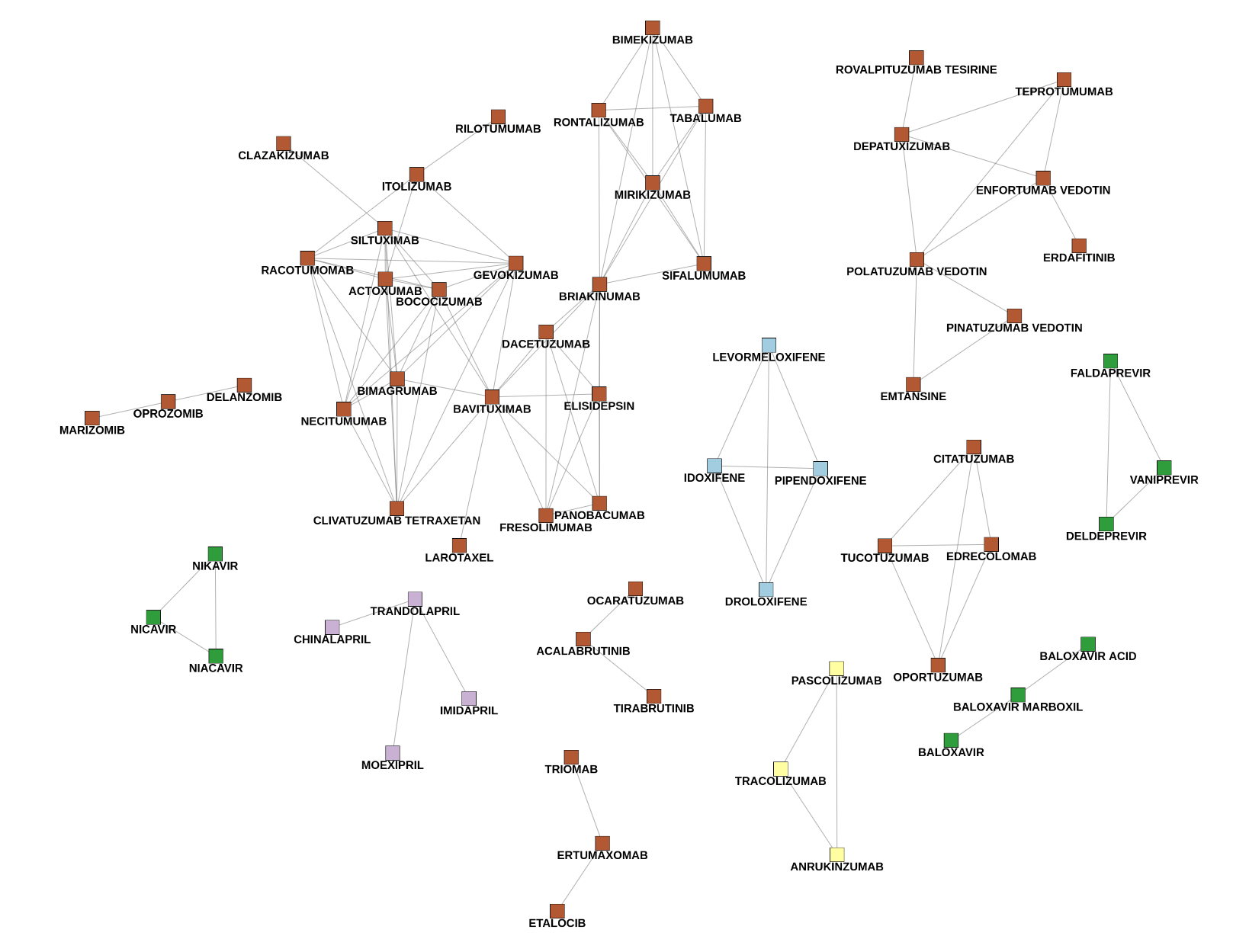

Fig. 7: Connected components consisting of unknown drugs only. The color describes different types of drugs based on their predicted ATC code. Each of these components will be extended with their neighbors from the complete network to evaluate the ATC predictions.
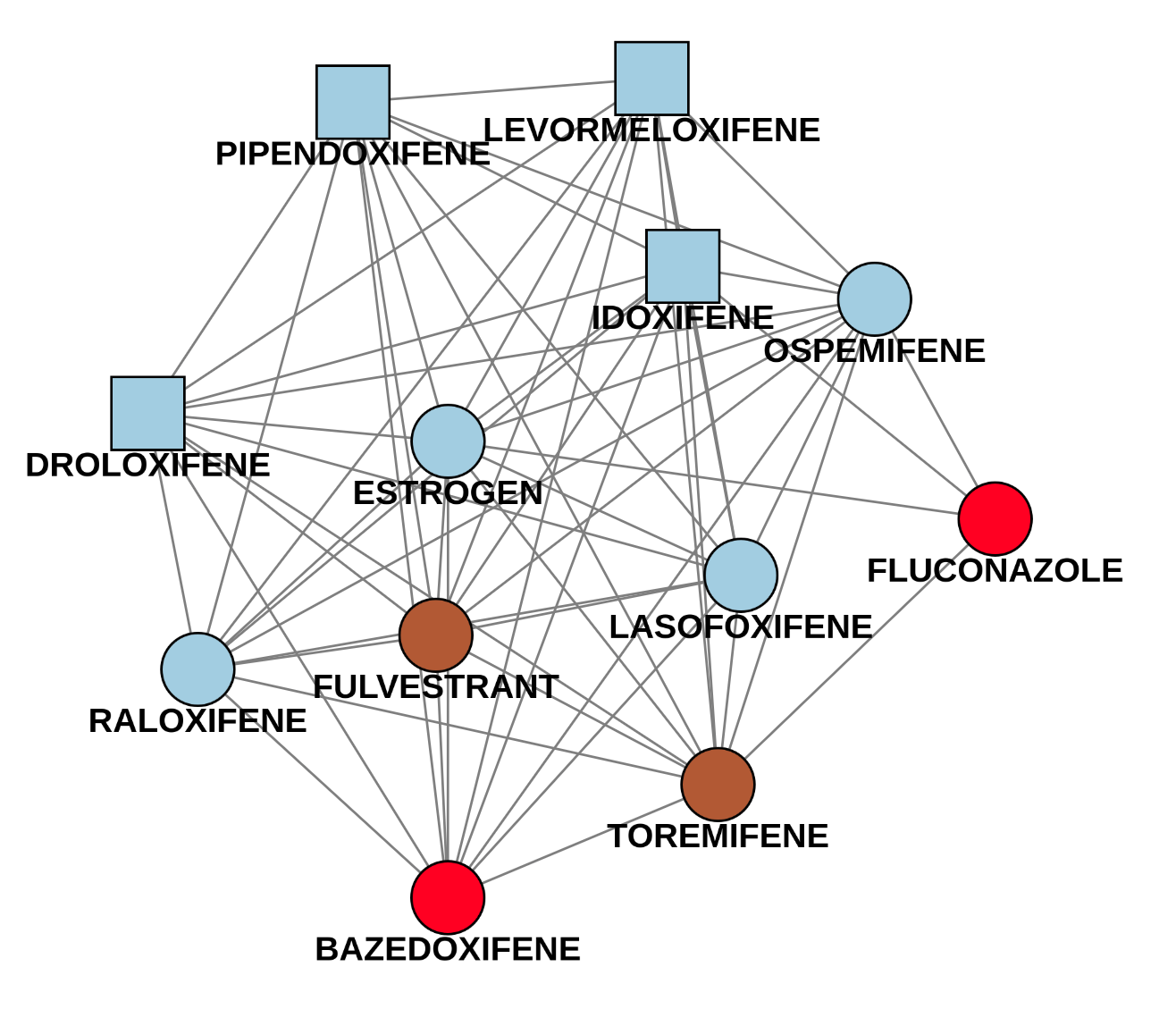
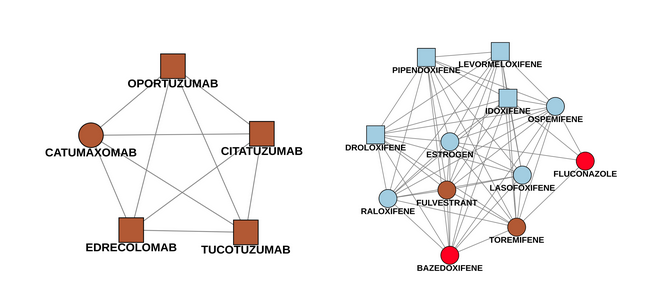
The following example (Fig.8) shows two of these subgraphs. The first picture is an easy case, since the four newly identified drug names only have one connection to a known drug (catumaxomab). All drugs were labeled as Antineoplastic and immunomodulating agents which is indeed correct. The second component is trickier. There are four newly detected drugs pipendoxifene, levormeloxifene, idoxifene and droloxifene. All of them were predicted as Genito-urinary system and sex hormones, since most of the known drugs in the network are in this ATC class (bazedoxifene included - it’s colored red because it has multiple ATC classes). However, there are also connections to Antineoplastic and immunomodulating agents like fulvestrant and toremifene. Connections to both of these drugs are worth mentioning as well, since the new drugs were mostly developed for breast cancer treatments. As we can see, the prediction might be right, but having a look at connections to ATC classes with a lower influence is also helpful to understand the purpose in a better way.
Summary
Today, we successfully trained a named-entity recognition model to detect drug names in biomedical literature and predict the purpose of the newly identified drugs. We started with an initial set of drug names from the World Health Organization, which also provides some more information about the drug’s purpose as they are annotated using the ATC Classification System. Based on this list, we then created a text corpus of articles by fetching them from PubMed. The StanfordNLP NE nodes then helped to train a named-entity recognition model to detect not only known drug names, but also some that were not in our initial data. Finally, we built a drug co-occurrence network to predict the purpose of unknown drugs based on their neighborhood and showed how to extract interesting subgraphs to easily evaluate our predictions.
The trained model and the prediction process can now be applied to any new literature, to get an instant overview of all drugs mentioned.
References
1. "Drug name recognition and classification in biomedical texts. A case ...."17 July 2008. Accessed 12 September 2019
2. "ATC/DDD Index - WHOCC"13 December 2018. Accessed 12 September 2019
The workflow group, Prediction of Drug Purpose, used for this blog post is available on the KNIME Hub under 08_Other_Analytics_Types/02_Chemistry_and_Life_Sciences/04_Prediction_Of_Drug_Purpose/
